
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 8,920,058 – 8,920,156 |
| Length | 98 |
| Max. P | 0.873848 |

| Location | 8,920,058 – 8,920,156 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 73.58 |
| Mean single sequence MFE | -25.98 |
| Consensus MFE | -15.39 |
| Energy contribution | -16.23 |
| Covariance contribution | 0.83 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.59 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.873848 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
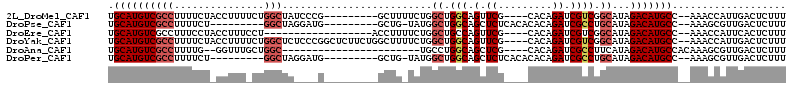
>2L_DroMel_CAF1 8920058 98 - 22407834 UGCAUGUCGCCUUUUCUACCUUUUCUGGCUAUCCCG---------GCUUUUCUGGCUGGCAGUUCG----CACAGAUCGUCGGCAUAGACAUGCC--AAACCAUUGACUCUUU .((((((((((...............)))(((.(((---------((...((((....((.....)----).))))..))))).)))))))))).--................ ( -28.56) >DroPse_CAF1 282638 92 - 1 UGCAUGUCGCCUUUUCU---------GGCUAGGAUG---------GCUG-UAUGGCUGGCAGCUCUCACACACAGAUCGCCUGCAUAGACAUGCC--AAAGCGUUGACUCUUU .((((((((((......---------)))...((.(---------((((-(.......)))))).)).....(((.....)))....))))))).--................ ( -26.10) >DroEre_CAF1 137915 89 - 1 UGCAUGUCGCCUUUCCUACCUUUCCU------------------ACCUUUUCUGGCUGCCAGUUCG----CACAGAUCGUCGGCAUAGACAUGCC--AAACCAUUCACUCUUU .((((((((((...............------------------.........)))((((.(.((.----....)).)...))))..))))))).--................ ( -19.56) >DroYak_CAF1 134496 107 - 1 UGCAUGUCGCCUUUUCUACCUUUUCUGGCUCUCCCGGCUCUUCUGGCUUUUCUGGCUGGCAGUUCG----CACAGAUCGUCGGCAUAGACAUGCC--AAACCAUUGACUCUUU .((((((((((...............)))....(((((...((((((....(((.....)))...)----).))))..)))))....))))))).--................ ( -30.26) >DroAna_CAF1 132637 84 - 1 UGCAUGUCGCCUUUUG--GGUUUGCUGGC-----------------------UGCCUGGCAGCUCG----CACAGAUCGCCUUCAUAGACAUGCCACAAAGCGUUGACUCUUU .((((((((((.....--))).(((.(((-----------------------(((...)))))).)----))...............)))))))................... ( -25.30) >DroPer_CAF1 186201 92 - 1 UGCAUGUCGCCUUUUCU---------GGCUAGGAUG---------GCUG-UAUGGCUGGCAGCUCUCACACACAGAUCGCCUGCAUAGACAUGCC--AAAGCGUUGACUCUUU .((((((((((......---------)))...((.(---------((((-(.......)))))).)).....(((.....)))....))))))).--................ ( -26.10) >consensus UGCAUGUCGCCUUUUCU_CCUUU_CUGGCUA____G_________GCUU_UCUGGCUGGCAGCUCG____CACAGAUCGCCGGCAUAGACAUGCC__AAACCAUUGACUCUUU .((((((((((...............))).........................((.(((.(.((.........)).)))).))...)))))))................... (-15.39 = -16.23 + 0.83)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:45:05 2006