
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 8,826,473 – 8,826,588 |
| Length | 115 |
| Max. P | 0.507994 |

| Location | 8,826,473 – 8,826,588 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 79.63 |
| Mean single sequence MFE | -45.00 |
| Consensus MFE | -30.87 |
| Energy contribution | -30.48 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.28 |
| Mean z-score | -1.13 |
| Structure conservation index | 0.69 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.507994 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
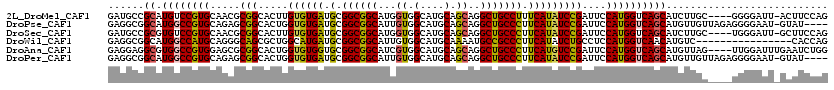
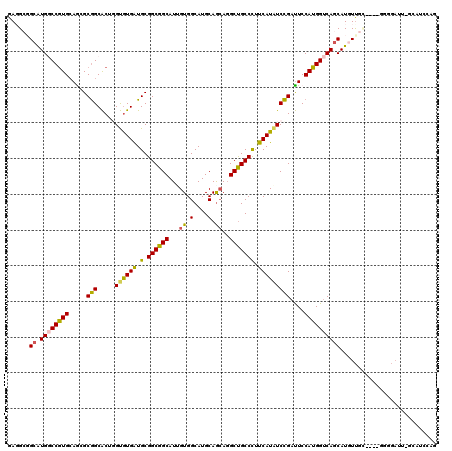
>2L_DroMel_CAF1 8826473 115 - 22407834 GAUGCCGCAUGUCCGUGCAACGCGGCACUUGUGUGAUGCGGCGGCAUGGUGGCAUGCAGCAGGCUGCCUUUCAUAUCCGAUUCCAUGGUCAGCAUCUUGC----GGGGAUU-ACUUCCAG ....((((((((((((((..((((.(((....))).))))...))))))..)))))).(((((.(((...((((..........))))...))).)))))----))(((..-...))).. ( -44.30) >DroPse_CAF1 53350 115 - 1 GAGGCGGCAUGGCCGUGCAGAGCGGCACUGGUGUGAUGCGGCGGCAUUGUGGCAUGCAGCAGGCUGCCCUUCAUAUCCGAUUCCAUGGUCAGCAUGUUGUUAGAGGGGAAU-GUAU---- ..((((((...(((((.....))))).(((.(((.((((.(((....))).))))))).))))))))).(((...((..((..((((.....))))..))..))...))).-....---- ( -43.70) >DroSec_CAF1 36617 115 - 1 GAUGCCGCGUGUCCGUGCAACGCGGCACUUGUGUGAUGCGGCGGCAUGGUGGCAUGCAGCAGGCUGCCCUUCAUAUCCGAUUCCAUGGUCAGCAUCUUGC----UGGGAUU-GCUUCCAG .((((((((((.(((((...)))))))).(((.....))))))))))((.(((((.(((((((.(((...((((..........))))...))).)))))----)).)..)-))).)).. ( -44.20) >DroWil_CAF1 75483 104 - 1 GAGGCGGCAUGGCCAUGCAGGGCAGCGCUGGCAUGAUGCGGCGGCAUUGUGGCAUGCAAAAUGCCGCCCUUCAUAUCUGCCUCCAUGGUCAACAUGUC----------------CACCAG ..((.((((((((((((..(((((((....))((((.(.(((((((((.((.....)).)))))))))).))))...))))).))))))...)))).)----------------).)).. ( -46.80) >DroAna_CAF1 36694 116 - 1 GAGGAGGCGUGGCCGUGGAGCGCGGCACUGGUGUGGUGCGGCGGCAUCGUGGCAUGCAGCAGGCUGCCCUUCAUGUCCGAUUCCAUGGUCAGCAUGUUAG----UUGGAUUUGAAUCUGG ..((((((...)))((((((.(((((.(((.((((.(((.(((....))).))))))).)))))))).)))))).)))(((((.....(((((......)----))))....)))))... ( -47.30) >DroPer_CAF1 54173 115 - 1 GAGGCGGCAUGGCCGUGCAGAGCGGCACUGGUGUGAUGCGGCGGCAUUGUGGCAUGCAGCAGGCUGCCCUUCAUAUCCGAUUCCAUGGUCAGCAUGUUGUUAGAGGGGAAU-GUAU---- ..((((((...(((((.....))))).(((.(((.((((.(((....))).))))))).))))))))).(((...((..((..((((.....))))..))..))...))).-....---- ( -43.70) >consensus GAGGCGGCAUGGCCGUGCAGCGCGGCACUGGUGUGAUGCGGCGGCAUUGUGGCAUGCAGCAGGCUGCCCUUCAUAUCCGAUUCCAUGGUCAGCAUGUUGC____GGGGAUU_GCAUCCAG ......((.((((((((.....(((.....((((((.(.((((((...((.(....).))..))))))).)))))))))....))))))))))........................... (-30.87 = -30.48 + -0.38)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:44:31 2006