| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 889,502 – 889,594 |
| Length | 92 |
| Max. P | 0.907683 |

| Location | 889,502 – 889,594 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 78.95 |
| Mean single sequence MFE | -18.70 |
| Consensus MFE | -12.10 |
| Energy contribution | -12.10 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.65 |
| SVM decision value | 1.05 |
| SVM RNA-class probability | 0.907683 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 889502 92 + 22407834 -CGUCCA-GUCCGCAGCCACAUUCAAAAUGCCAUUAAAAGCAUACUAAAAUUAAUUUGUAGCAAUUAGUUGCAAAUUGCCAGA--AAGCAAGGCGG--------------------- -......-..((((.((....(((...((((........))))........(((((((((((.....)))))))))))...))--).))...))))--------------------- ( -19.70) >DroGri_CAF1 20298 102 + 1 -CGUCGACGUCAUUUACCACAUUCAAAAUGCCAUUAAAAGCAUACUAAAAUUAAUUUGUAGCAAUUAGUUGCAAAUUGCAAGG--AAGCGAAAUGAAAAU-----GAGAA------- -..(((...((((((......(((...((((........))))........(((((((((((.....)))))))))))....)--))...))))))...)-----))...------- ( -18.80) >DroEre_CAF1 21184 95 + 1 -CGUCCA-GUCCGCAGCCACAUUCAAAAUGCCAUUAAAAGCAUACUAAAAUUAAUUUGUAGCAAUUAGUUGCAAAUUGCCAGA--AAGCAAGGCAGCCG------------------ -......-....((.(((.........((((........))))........(((((((((((.....))))))))))).....--......))).))..------------------ ( -19.50) >DroWil_CAF1 26344 91 + 1 CCCU-CA-U--UUAUACCACAUUCAAAAUGCCAUUAAAAGCAUACUAAAAUUAAUUUGUAGCAAUUAGUUGCAAAUUGCAUGAAAAAGCAAACCA---------------------- ....-..-.--..........((((..((((........))))........(((((((((((.....)))))))))))..))))...........---------------------- ( -14.10) >DroYak_CAF1 17769 113 + 1 -CGUCCA-GUCGGCAGCCACAUUCAAAAUGCCAUUAAAAGCAUACUAAAAUUAAUUUGUAGCAAUUAGUUGCAAAUUGCCAGA--AAGCAAGGCGGCAGUCAAAAGCAAAAGCAAAA -......-...(((.(((.........((((........)))).........((((((((((.....))))))))))(((...--......)))))).)))....((....)).... ( -24.60) >DroAna_CAF1 19099 90 + 1 -CGGCCC-GUCC-CAACCACAUUCAAAAUGCCAUUAAAAGCAUACUAAAAUUAAUUUGUAGCAAUUAGUUGCAAAUUGCAAGA--AAGCACAGCC---------------------- -.(((..-....-..............((((........))))........(((((((((((.....))))))))))).....--.......)))---------------------- ( -15.50) >consensus _CGUCCA_GUCCGCAACCACAUUCAAAAUGCCAUUAAAAGCAUACUAAAAUUAAUUUGUAGCAAUUAGUUGCAAAUUGCAAGA__AAGCAAAGCGG_____________________ ...........................((((........))))........(((((((((((.....)))))))))))....................................... (-12.10 = -12.10 + -0.00)

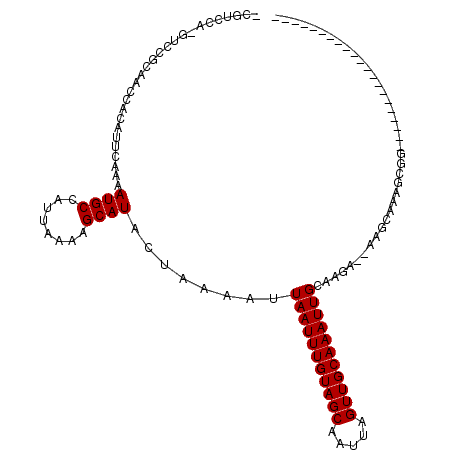
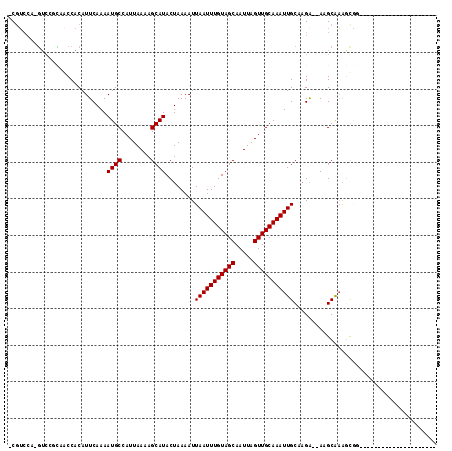
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:32:38 2006