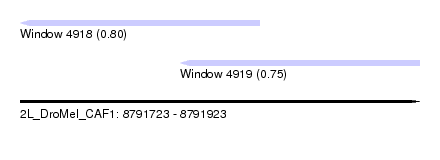
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 8,791,723 – 8,791,923 |
| Length | 200 |
| Max. P | 0.796282 |

| Location | 8,791,723 – 8,791,843 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.89 |
| Mean single sequence MFE | -42.50 |
| Consensus MFE | -41.57 |
| Energy contribution | -41.90 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.98 |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.796282 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 8791723 120 - 22407834 CAUGCUGGCCAUGGAUUCCAUGGUUGGGGGACCGAGAAUUGAGGACUGGCAGGACUCUGUUAUUUUAAUGAAAAUUGGAAAGUGCUCCUCGCUCCGCUUGGCCCCAGUUCCUGCUCCAAU ...((.((((((((...)))))))).((((.(((((...((((((.((((((....))))))(((((((....))))))).....)))))).....))))))))).......))...... ( -41.90) >DroSec_CAF1 94413 120 - 1 CAUGCUGGCCAUGGAUUCCAUGGUUUGGGGACCGAGAAUUGAGGACUGGCAGGACUCUGUUAUUUUAAUGAAAAUUGGAAAGUGCUCCUCGCUCCGCUUGGCCCCAGUUCCUGCUCCAAU ...((.((((((((...)))))).((((((.(((((...((((((.((((((....))))))(((((((....))))))).....)))))).....)))))))))))..)).))...... ( -42.80) >DroSim_CAF1 97991 120 - 1 CAUGCUGGCCAUGGGUUCCAUGGUUUGGGGACCGAGAAUUGAGGACUGGCAGGACUCUGUUAUUUUAAUGAAAAUUGGAAAGUGCUCCUCGCUCCGCUUGGCCCCAGUUCCUGCUCCAAU ...((.((((((((...)))))).((((((.(((((...((((((.((((((....))))))(((((((....))))))).....)))))).....)))))))))))..)).))...... ( -42.80) >consensus CAUGCUGGCCAUGGAUUCCAUGGUUUGGGGACCGAGAAUUGAGGACUGGCAGGACUCUGUUAUUUUAAUGAAAAUUGGAAAGUGCUCCUCGCUCCGCUUGGCCCCAGUUCCUGCUCCAAU ...((.((((((((...)))))).((((((.(((((...((((((.((((((....))))))(((((((....))))))).....)))))).....)))))))))))..)).))...... (-41.57 = -41.90 + 0.33)

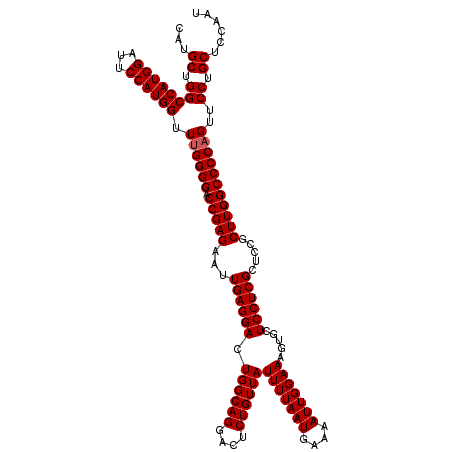
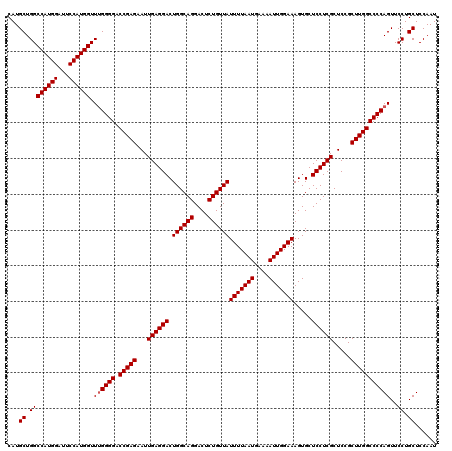
| Location | 8,791,803 – 8,791,923 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.21 |
| Mean single sequence MFE | -38.10 |
| Consensus MFE | -34.14 |
| Energy contribution | -33.70 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.49 |
| SVM RNA-class probability | 0.754727 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 8791803 120 - 22407834 CCCCAUCUGCAUCCCCCAACUCAACCCCUCAAAAACCUGCGCCCUAAAAAGUACUCUUCUGGGGGUGGCUAUCCAAUCGCCAUGCUGGCCAUGGAUUCCAUGGUUGGGGGACCGAGAAUU .....(((...(((((((((.(((((((.((......(((..........)))......)))))))((..(((((...(((.....)))..))))).)).)))))))))))...)))... ( -41.10) >DroSec_CAF1 94493 119 - 1 CCCCAUCUGCAUCCCCCAACUCAACCCC-GAAAAACCUGCGCCCUAAAAAGUACUCUUCUGGGGGUGGCUAUCCAAUCGCCAUGCUGGCCAUGGAUUCCAUGGUUUGGGGACCGAGAAUU ...................(((..((((-(((...((.((.(((((..(((....))).)))))(((((.........))))))).))((((((...)))))))))))))...))).... ( -36.60) >DroSim_CAF1 98071 119 - 1 CCCCAUCUGCAUCCCCCCACUCAACCCC-GAAAAACCUGCGCCCUAAAAAGUACUCUUCUGGGGGUGGCUAUCCAAUCGCCAUGCUGGCCAUGGGUUCCAUGGUUUGGGGACCGAGAAUU ...................(((..((((-(((...((.((.(((((..(((....))).)))))(((((.........))))))).))((((((...)))))))))))))...))).... ( -36.60) >consensus CCCCAUCUGCAUCCCCCAACUCAACCCC_GAAAAACCUGCGCCCUAAAAAGUACUCUUCUGGGGGUGGCUAUCCAAUCGCCAUGCUGGCCAUGGAUUCCAUGGUUUGGGGACCGAGAAUU ...................(((..((((.(((...((.((.(((((..(((....))).)))))(((((.........))))))).))((((((...)))))))))))))...))).... (-34.14 = -33.70 + -0.44)

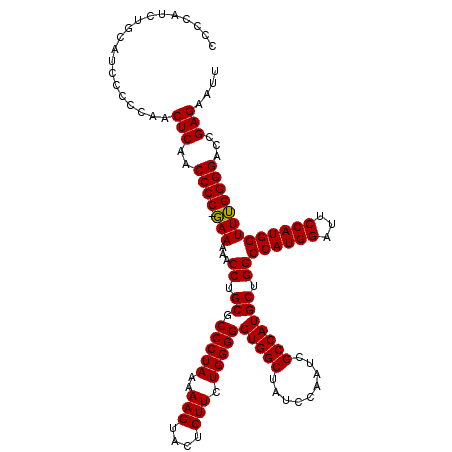
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:44:08 2006