
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 8,630,828 – 8,630,920 |
| Length | 92 |
| Max. P | 0.820719 |

| Location | 8,630,828 – 8,630,920 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 76.58 |
| Mean single sequence MFE | -19.04 |
| Consensus MFE | -9.92 |
| Energy contribution | -10.42 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.17 |
| SVM RNA-class probability | 0.616098 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 8630828 92 + 22407834 GAUAAUCCGCUUGCCAUU---UUCCUUUCUUC-UCAACUGCCUUUUGUCGCCUUUAACGCCGCCUAAUUGCCUAAUUAGGCAUUUGCUGGCCGAAC (((((...((........---...........-......))...)))))(((..(((....((((((((....))))))))..)))..)))..... ( -16.01) >DroVir_CAF1 77682 74 + 1 ---------GUUGGCAA------CCCUUAGCCGUGACU---UUUUUGUCGCCUUUAAUGACGCCUAAUUGCCUAAUUAGGCAUUUGCUGG----CC ---------((..((((------.(.((((..(((((.---.....)))))..)))).)..((((((((....))))))))..))))..)----). ( -24.70) >DroSim_CAF1 65330 92 + 1 GAUAAUCCGCUUGCCAUU---UUCCUUUCUUC-UCAACUGCCUUUUGUCGCCUUUAACGCCGCCUAAUUGCCUAAUUAGGCAUUUGCUGGCCGAAC (((((...((........---...........-......))...)))))(((..(((....((((((((....))))))))..)))..)))..... ( -16.01) >DroEre_CAF1 60356 95 + 1 GAUAAUCCGCUUGCCUUUUUUUUGCUUUCUUC-UCAACUGCCUUUUGUCGCCUUUAACGCCGCCUAAUUGCCUAAUUAGGCAUUUGAUGGCCGAAC (((((...((......................-......))...)))))(((.((((....((((((((....))))))))..)))).)))..... ( -16.99) >DroYak_CAF1 61978 93 + 1 GAUAAUCCGCUUGCGUUUU--UUCCAUUCUUC-UCAACUGCCUUUUGUCGCCUUUAACGCCGCCUAAUUGCCUAAUUAGGCAUUUGCUGGCCGAAC .....((.(((.(((....--...........-......((........))..........((((((((....))))))))...))).))).)).. ( -16.60) >DroMoj_CAF1 89305 80 + 1 ---------CUGGCCAA------CCCUUAGCC-UGGCUUUUUUUUUGUCGCAUUUAACGAUGCCUAAUUGCCUAAUUAGGCAUUUGCAGCCAGAUC ---------(((((...------.........-.(((.........)))(((......(((((((((((....)))))))))))))).)))))... ( -23.90) >consensus GAUAAUCCGCUUGCCAUU___UUCCUUUCUUC_UCAACUGCCUUUUGUCGCCUUUAACGCCGCCUAAUUGCCUAAUUAGGCAUUUGCUGGCCGAAC .................................................(((..(((....((((((((....))))))))..)))..)))..... ( -9.92 = -10.42 + 0.50)


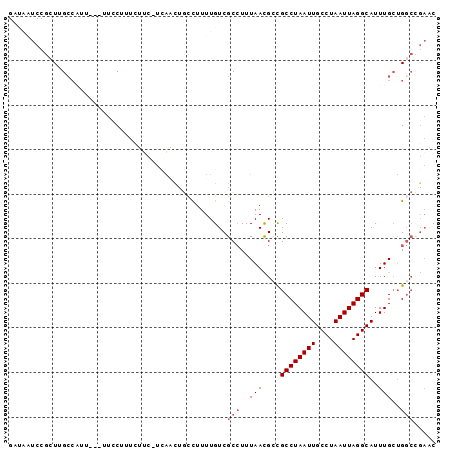
| Location | 8,630,828 – 8,630,920 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 76.58 |
| Mean single sequence MFE | -22.75 |
| Consensus MFE | -11.23 |
| Energy contribution | -11.59 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.49 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.820719 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 8630828 92 - 22407834 GUUCGGCCAGCAAAUGCCUAAUUAGGCAAUUAGGCGGCGUUAAAGGCGACAAAAGGCAGUUGA-GAAGAAAGGAA---AAUGGCAAGCGGAUUAUC (((((((((((....((((((((....)))))))).)).((((..((........))..))))-...........---..))))...))))).... ( -21.90) >DroVir_CAF1 77682 74 - 1 GG----CCAGCAAAUGCCUAAUUAGGCAAUUAGGCGUCAUUAAAGGCGACAAAAA---AGUCACGGCUAAGGG------UUGCCAAC--------- ((----(.(((...(((((....))))).((((.((((......)))(((.....---.)))..).))))..)------)))))...--------- ( -22.50) >DroSim_CAF1 65330 92 - 1 GUUCGGCCAGCAAAUGCCUAAUUAGGCAAUUAGGCGGCGUUAAAGGCGACAAAAGGCAGUUGA-GAAGAAAGGAA---AAUGGCAAGCGGAUUAUC (((((((((((....((((((((....)))))))).)).((((..((........))..))))-...........---..))))...))))).... ( -21.90) >DroEre_CAF1 60356 95 - 1 GUUCGGCCAUCAAAUGCCUAAUUAGGCAAUUAGGCGGCGUUAAAGGCGACAAAAGGCAGUUGA-GAAGAAAGCAAAAAAAAGGCAAGCGGAUUAUC ((((((((......(((((((((....)))))))))((........((((........)))).-.......))........)))...))))).... ( -21.79) >DroYak_CAF1 61978 93 - 1 GUUCGGCCAGCAAAUGCCUAAUUAGGCAAUUAGGCGGCGUUAAAGGCGACAAAAGGCAGUUGA-GAAGAAUGGAA--AAAACGCAAGCGGAUUAUC ..((.((..((...(((((((((....))))))))).((((.....((((........)))).-....))))...--.....))..)).))..... ( -21.80) >DroMoj_CAF1 89305 80 - 1 GAUCUGGCUGCAAAUGCCUAAUUAGGCAAUUAGGCAUCGUUAAAUGCGACAAAAAAAAAGCCA-GGCUAAGGG------UUGGCCAG--------- ....(((((.....(((((((((....)))))))))((((.....)))).........)))))-(((((....------.)))))..--------- ( -26.60) >consensus GUUCGGCCAGCAAAUGCCUAAUUAGGCAAUUAGGCGGCGUUAAAGGCGACAAAAGGCAGUUGA_GAAGAAAGGAA___AAUGGCAAGCGGAUUAUC .....(((......(((((((((....)))))))))........)))................................................. (-11.23 = -11.59 + 0.36)


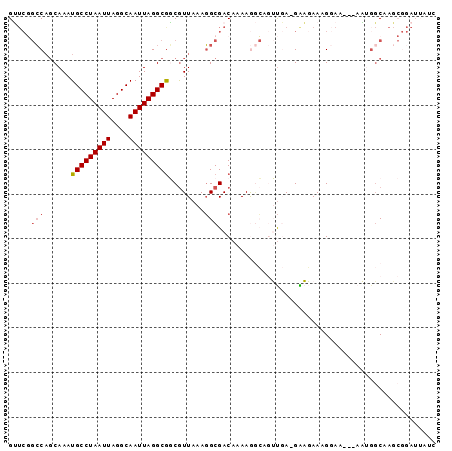
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:42:39 2006