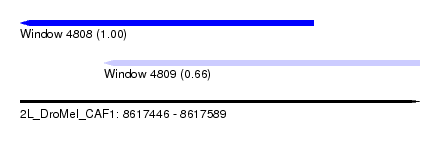

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 8,617,446 – 8,617,589 |
| Length | 143 |
| Max. P | 0.995487 |

| Location | 8,617,446 – 8,617,551 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 96.36 |
| Mean single sequence MFE | -18.71 |
| Consensus MFE | -16.70 |
| Energy contribution | -17.30 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.89 |
| SVM decision value | 2.58 |
| SVM RNA-class probability | 0.995487 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
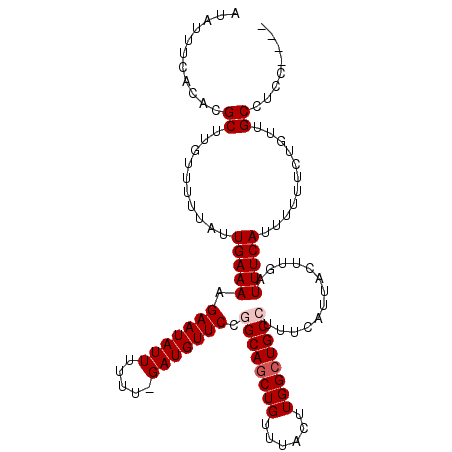
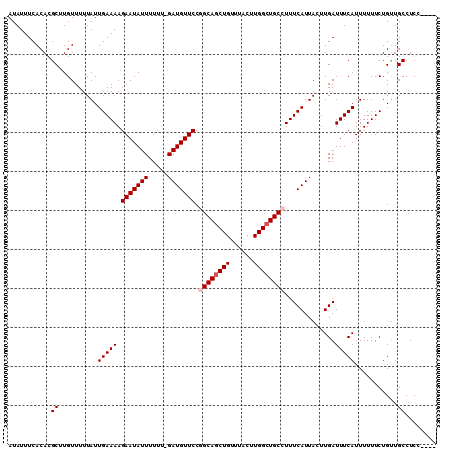
>2L_DroMel_CAF1 8617446 105 - 22407834 AUAUUUCACACGCUUGUUUUUAUUGAAAAGAAUAUUUUUU-GAUGUUCCGGCAGCUGUUUACUUGGCUGCCUUUCAUUACUUGAUUUCAUUUUUUCUGUUGCCUCC---- ...........((..........(((((.(((((((....-))))))).((((((((......)))))))).............)))))...........))....---- ( -21.30) >DroSec_CAF1 45744 105 - 1 AUAUUUCACACGCUUGUUUUUAUUGAAAAGAAUAUUUUUU-GAUGUUCCUGCAGCUGUUUACUUGGCUGCCUUUCAUUAUUUGAUUUCAUUUUUUCUGUUGCCUCC---- ...........((..........(((((.(((((((....-)))))))..(((((((......)))))))..............)))))...........))....---- ( -17.30) >DroSim_CAF1 51430 105 - 1 AUAUUUCACACGCUUGUUUUUAUUGAAAAGAAUAUUUUUU-GAUGUUCCAGCAGCUGUUUACUUGGCUGCCUUUCAUUACUUGAUUUCAUUUUUUCUGUUGCCUCC---- ...........((..........(((((.(((((((....-)))))))..(((((((......)))))))..............)))))...........))....---- ( -17.50) >DroEre_CAF1 47395 105 - 1 AUAUUUCACACGCUUGUUUUUAUUGAAAAGAAUAUUUUUU-GAUGUUCCGGCAGCUGUUUACUUGGCUGCCUUUCAUUACUUGAUUUCAUUUUUUCUGCUGCCUCC---- ...........((..((......(((((.(((((((....-))))))).((((((((......)))))))).............)))))........)).))....---- ( -21.84) >DroYak_CAF1 48688 110 - 1 AUAUUUCACACGCUUGUUUUUAUUGAAAAGAAUAUUUUUUCGAUGUUCCGGCAACUGUUUACUUGGCUGCCUUUCAUUACUUGAUUUCAUUUUUUCUGUUGCCUCCAUUC ....................(((((((((((...)))))))))))....((((((.(((.....)))......(((.....))).............))))))....... ( -15.60) >consensus AUAUUUCACACGCUUGUUUUUAUUGAAAAGAAUAUUUUUU_GAUGUUCCGGCAGCUGUUUACUUGGCUGCCUUUCAUUACUUGAUUUCAUUUUUUCUGUUGCCUCC____ ...........((..........(((((.(((((((.....))))))).((((((((......)))))))).............)))))...........))........ (-16.70 = -17.30 + 0.60)

| Location | 8,617,476 – 8,617,589 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 93.48 |
| Mean single sequence MFE | -26.80 |
| Consensus MFE | -20.48 |
| Energy contribution | -22.24 |
| Covariance contribution | 1.76 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.661073 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
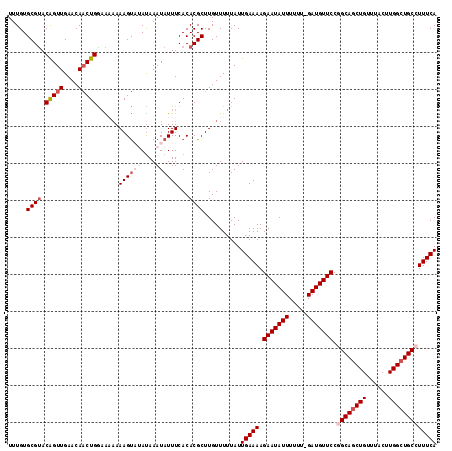
>2L_DroMel_CAF1 8617476 113 - 22407834 UUUGUGCGUACAGUUGAACAACUGGAA--AAAGUAUAUAAAUAUUUCACACGCUUGUUUUUAUUGAAAAGAAUAUUUUUU-GAUGUUCCGGCAGCUGUUUACUUGGCUGCCUUUCA .....((((.(((((....)))))...--.((((((....))))))...))))..........(((((.(((((((....-))))))).((((((((......))))))))))))) ( -29.30) >DroSec_CAF1 45774 115 - 1 UUUGUGCGUACAGUUGAACAACUGAAAAAAAAGUAUAUAAAUAUUUCACACGCUUGUUUUUAUUGAAAAGAAUAUUUUUU-GAUGUUCCUGCAGCUGUUUACUUGGCUGCCUUUCA .....((((.(((((....)))))......((((((....))))))...))))..........(((((.(((((((....-)))))))..(((((((......))))))).))))) ( -25.10) >DroSim_CAF1 51460 114 - 1 UUUGUGCGUACAGUUGAACAACUGAAA-AAAAGUAUAUAAAUAUUUCACACGCUUGUUUUUAUUGAAAAGAAUAUUUUUU-GAUGUUCCAGCAGCUGUUUACUUGGCUGCCUUUCA .....((((.(((((....)))))...-..((((((....))))))...))))..........(((((.(((((((....-)))))))..(((((((......))))))).))))) ( -25.30) >DroEre_CAF1 47425 112 - 1 UUUGUGCGCACGGUUGAACA-CUGGAGAAAAAGC--ACAAAUAUUUCACACGCUUGUUUUUAUUGAAAAGAAUAUUUUUU-GAUGUUCCGGCAGCUGUUUACUUGGCUGCCUUUCA (((((((.(.((((.....)-)))..).....))--)))))......................(((((.(((((((....-))))))).((((((((......))))))))))))) ( -28.70) >DroYak_CAF1 48722 114 - 1 UUUGUGCGUACGGUUGAACAACUGGUGAAAAAGU--AUAAAUAUUUCACACGCUUGUUUUUAUUGAAAAGAAUAUUUUUUCGAUGUUCCGGCAACUGUUUACUUGGCUGCCUUUCA .....((((.(((((....)))))((((((....--.......)))))))))).......(((((((((((...)))))))))))....((((.(((......))).))))..... ( -25.60) >consensus UUUGUGCGUACAGUUGAACAACUGGAAAAAAAGUAUAUAAAUAUUUCACACGCUUGUUUUUAUUGAAAAGAAUAUUUUUU_GAUGUUCCGGCAGCUGUUUACUUGGCUGCCUUUCA .....((((.(((((....)))))......((((((....))))))...))))..........(((((.(((((((.....))))))).((((((((......))))))))))))) (-20.48 = -22.24 + 1.76)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:42:24 2006