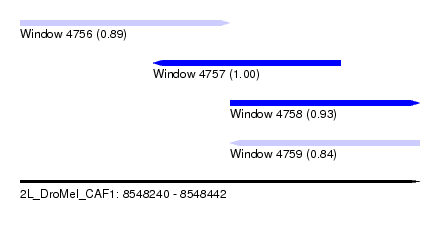

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 8,548,240 – 8,548,442 |
| Length | 202 |
| Max. P | 0.995623 |

| Location | 8,548,240 – 8,548,346 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 94.19 |
| Mean single sequence MFE | -19.84 |
| Consensus MFE | -16.56 |
| Energy contribution | -15.96 |
| Covariance contribution | -0.60 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.98 |
| SVM RNA-class probability | 0.893405 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
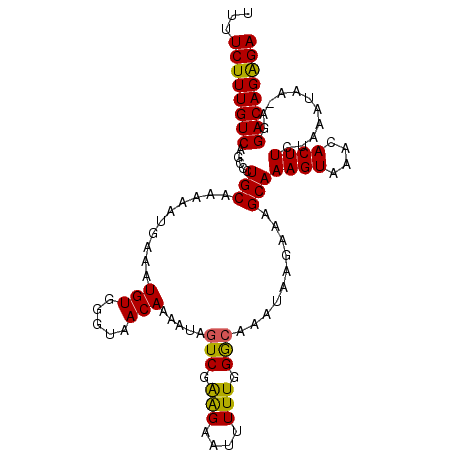
>2L_DroMel_CAF1 8548240 106 + 22407834 UUUUCUUUGUCAGGCCUGCAAAAAUGAAAUGUGAGUAACAAAAUAGUCGAAGAAUUUUUGGGCAAAUAAGAAAGCAAAGUAAACACUUCUAAAUAA-AGGACAGAGA ...((((((((..(((..(((((((......(((.((......)).)))....)))))))))).............((((....))))........-..)))))))) ( -19.80) >DroSec_CAF1 57766 106 + 1 UUUUCUUUGUCAGCCCUGCAAAAAUGAAAUGUGGGUAACAAAAUAGUCGAGGAAUUUUUGGACAAAUAAGAAAGCAAAGUAAACACUUCUAAAUAA-AGGACAGAGA ...((((((((.((((..((.........))..))..........(((.(((....))).)))..........)).((((....))))........-..)))))))) ( -19.30) >DroSim_CAF1 57478 106 + 1 UUUUCUUUGUCAGCCCUGCAAAAAUGAAAUGUGGGUAACAAAAUAGUCGAGGAAUUUUUGGACAAAUAAGAAAGCAAAGUAAACACUUCUAAAUAA-AGGACAGAGA ...((((((((.((((..((.........))..))..........(((.(((....))).)))..........)).((((....))))........-..)))))))) ( -19.30) >DroEre_CAF1 59276 107 + 1 UUUUCUUUGUCAGCCCUGCAAAAAUGAAAUGUGGGUAACAAAAUAGUCGGAGAAUUUUUGGGAAAACAAGAAAGCAAAGUAAACACUUAUAGAUAAAAGGACAGGGA ...((((((((.((((.(((.........))))))).........(((...(..((((((......))))))..).((((....))))...))).....)))))))) ( -20.80) >DroYak_CAF1 58883 107 + 1 UUUUCUUUGUCAGCCCUGCAAAAAUGAAAUGUGGGUAACAAAAAAGUCGAAGAAUUUUUGGGCAAACAGGAAAGCAAAGUAAACACUUAUAAAUAAAAGGACAGAGA ...((((((((.((((((...........(((.....))).....(((.(((....))).)))...))))...)).((((....))))...........)))))))) ( -20.00) >consensus UUUUCUUUGUCAGCCCUGCAAAAAUGAAAUGUGGGUAACAAAAUAGUCGAAGAAUUUUUGGGCAAAUAAGAAAGCAAAGUAAACACUUCUAAAUAA_AGGACAGAGA ...((((((((.....(((..........(((.....))).....(((.(((....))).)))..........)))((((....))))...........)))))))) (-16.56 = -15.96 + -0.60)


| Location | 8,548,307 – 8,548,402 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 92.07 |
| Mean single sequence MFE | -20.29 |
| Consensus MFE | -16.88 |
| Energy contribution | -17.92 |
| Covariance contribution | 1.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.83 |
| SVM decision value | 2.60 |
| SVM RNA-class probability | 0.995623 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
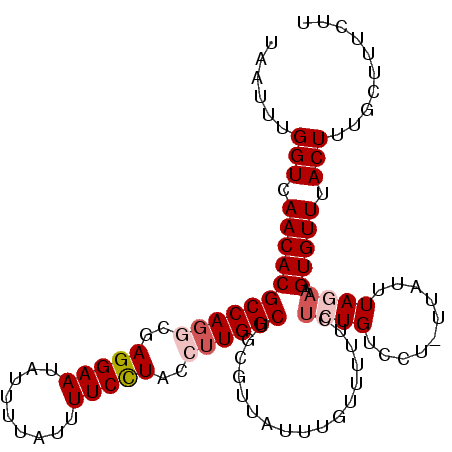
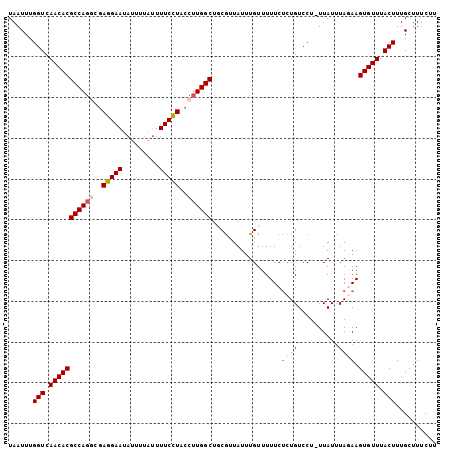
>2L_DroMel_CAF1 8548307 95 - 22407834 UAAUUUGGUCAACACGCCA----AGGAAUAU-----UUUCCUACCUUGGCUGCGUUAUUUGUUUUUCUCUGUCCU-UUAUUUAGAAGUGUUUACUUUGCUUUCUU ......(((.(((((((((----(((.....-----.......))))))).................((((....-.....)))).))))).))).......... ( -18.70) >DroSec_CAF1 57833 104 - 1 UAAUUUGGUCAACACGCCAGGCGAGGAAUAUUUUAUUUUCCUACCUUGGCUGCGUUAUUUGUUUUUCUCUGUCCU-UUAUUUAGAAGUGUUUACUUUGCUUUCUU ......(((.(((((((((((..(((((.........)))))..)))))).................((((....-.....)))).))))).))).......... ( -22.30) >DroSim_CAF1 57545 104 - 1 UAAUUUGGUCAACACGCCAGGCGAGGAAUAUUUUAUUUUCCUACCUUGGCUGCAUUAUUUGUUUUUCUCUGUCCU-UUAUUUAGAAGUGUUUACUUUGCUUUCUU ......(((.(((((((((((..(((((.........)))))..)))))).................((((....-.....)))).))))).))).......... ( -22.30) >DroEre_CAF1 59343 105 - 1 UAAUUUGGUCAACACGCCAGGCGAGGAAUAUUUUAUUUUCCUACCUUGGCUUCGCUAUUUGUUUUUCCCUGUCCUUUUAUCUAUAAGUGUUUACUUUGCUUUCUU ......(((.(((((((((((..(((((.........)))))..))))))...((.....))........................))))).))).......... ( -20.50) >DroYak_CAF1 58950 105 - 1 UAAUUUGGUCAACACGCCAGCCGAGGAAUAUUUUAUUUUCUUACCUUGGCUUCGCUAUUUGUUUUUCUCUGUCCUUUUAUUUAUAAGUGUUUACUUUGCUUUCCU ......(((.(((((((.((((((((.................))))))))..)).....................(((....)))))))).))).......... ( -17.63) >consensus UAAUUUGGUCAACACGCCAGGCGAGGAAUAUUUUAUUUUCCUACCUUGGCUGCGUUAUUUGUUUUUCUCUGUCCU_UUAUUUAGAAGUGUUUACUUUGCUUUCUU ......(((.(((((((((((..(((((.........)))))..)))))).................((((..........)))).))))).))).......... (-16.88 = -17.92 + 1.04)

| Location | 8,548,346 – 8,548,442 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 89.58 |
| Mean single sequence MFE | -28.77 |
| Consensus MFE | -21.24 |
| Energy contribution | -22.64 |
| Covariance contribution | 1.40 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.58 |
| Structure conservation index | 0.74 |
| SVM decision value | 1.23 |
| SVM RNA-class probability | 0.934130 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
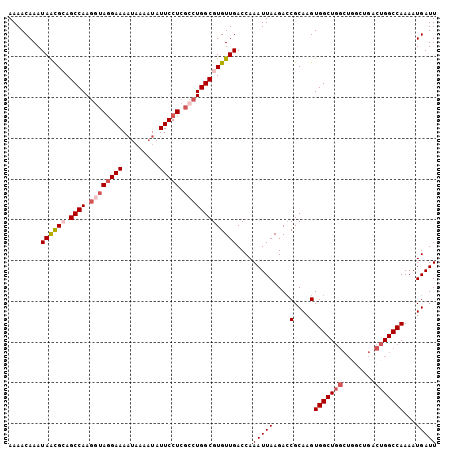
>2L_DroMel_CAF1 8548346 96 + 22407834 AAAACAAAUAACGCAGCCAAGGUAGGAAA-----AUAUUCCU----UGGCGUGUUGACCAAAUUAAGACCGCAAGUGGCUGGCUGGCUGACUGGCCAAAAUGAUU ........((((((.(((((((.......-----.....)))----))))))))))....(((((((.(((....)))))(((..(....)..)))....))))) ( -28.10) >DroSec_CAF1 57872 105 + 1 AAAACAAAUAACGCAGCCAAGGUAGGAAAAUAAAAUAUUCCUCGCCUGGCGUGUUGACCAAAUUAAGAGCGCAAGUGGCUGGCUGGCUGACUGGCCAAAAUGAUU ........((((((.((((.((((((((.........))))).)))))))))))))....(((((((.((....))..))(((..(....)..)))....))))) ( -31.60) >DroSim_CAF1 57584 101 + 1 AAAACAAAUAAUGCAGCCAAGGUAGGAAAAUAAAAUAUUCCUCGCCUGGCGUGUUGACCAAAUUAAGACCGCAAGUGGCU----GGCUGACUGGCCAAAAUGAUU ...........(((.((((.((((((((.........))))).)))))))((.((((.....)))).)).)))..((((.----.(....)..))))........ ( -28.50) >DroEre_CAF1 59383 101 + 1 AAAACAAAUAGCGAAGCCAAGGUAGGAAAAUAAAAUAUUCCUCGCCUGGCGUGUUGACCAAAUUAAGACCGCAAGUGGCUGGCU----GGCUGGCCAAAAUGAUU ..........(((..((((.((((((((.........))))).)))))))((.((((.....)))).)))))...((((..(..----..)..))))........ ( -30.80) >DroYak_CAF1 58990 101 + 1 AAAACAAAUAGCGAAGCCAAGGUAAGAAAAUAAAAUAUUCCUCGGCUGGCGUGUUGACCAAAUUAAGACCGCAAGUGGCUGGCU----GGCUGGCCAAAAUGAUU ....(((...(((.((((.(((.................))).))))..))).)))....(((((....(....)((((..(..----..)..))))...))))) ( -24.83) >consensus AAAACAAAUAACGCAGCCAAGGUAGGAAAAUAAAAUAUUCCUCGCCUGGCGUGUUGACCAAAUUAAGACCGCAAGUGGCUGGCUGGCUGACUGGCCAAAAUGAUU ........((((((.((((.((((((((.........))))).)))))))))))))....(((((....(....)(((((((........)))))))...))))) (-21.24 = -22.64 + 1.40)


| Location | 8,548,346 – 8,548,442 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 89.58 |
| Mean single sequence MFE | -25.39 |
| Consensus MFE | -20.39 |
| Energy contribution | -20.83 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.843470 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 8548346 96 - 22407834 AAUCAUUUUGGCCAGUCAGCCAGCCAGCCACUUGCGGUCUUAAUUUGGUCAACACGCCA----AGGAAUAU-----UUUCCUACCUUGGCUGCGUUAUUUGUUUU .........(((......)))(((((((((.....(((.......((((......))))----(((((...-----.)))))))).)))))).)))......... ( -23.90) >DroSec_CAF1 57872 105 - 1 AAUCAUUUUGGCCAGUCAGCCAGCCAGCCACUUGCGCUCUUAAUUUGGUCAACACGCCAGGCGAGGAAUAUUUUAUUUUCCUACCUUGGCUGCGUUAUUUGUUUU .......((((((((......((((((....))).)))......))))))))...((((((..(((((.........)))))..))))))............... ( -26.70) >DroSim_CAF1 57584 101 - 1 AAUCAUUUUGGCCAGUCAGCC----AGCCACUUGCGGUCUUAAUUUGGUCAACACGCCAGGCGAGGAAUAUUUUAUUUUCCUACCUUGGCUGCAUUAUUUGUUUU .......((((((((..((((----.((.....)))).))....))))))))...((((((..(((((.........)))))..))))))............... ( -26.30) >DroEre_CAF1 59383 101 - 1 AAUCAUUUUGGCCAGCC----AGCCAGCCACUUGCGGUCUUAAUUUGGUCAACACGCCAGGCGAGGAAUAUUUUAUUUUCCUACCUUGGCUUCGCUAUUUGUUUU ...((...((((.((((----((((.(((......))).......((((......))))))).(((((.........)))))....)))))..))))..)).... ( -25.90) >DroYak_CAF1 58990 101 - 1 AAUCAUUUUGGCCAGCC----AGCCAGCCACUUGCGGUCUUAAUUUGGUCAACACGCCAGCCGAGGAAUAUUUUAUUUUCUUACCUUGGCUUCGCUAUUUGUUUU .......((((((((..----((((.((.....)))).))....))))))))...((.((((((((.................))))))))..)).......... ( -24.13) >consensus AAUCAUUUUGGCCAGUCAGCCAGCCAGCCACUUGCGGUCUUAAUUUGGUCAACACGCCAGGCGAGGAAUAUUUUAUUUUCCUACCUUGGCUGCGUUAUUUGUUUU .......((((((((...........((.....)).........))))))))...((((((..(((((.........)))))..))))))............... (-20.39 = -20.83 + 0.44)

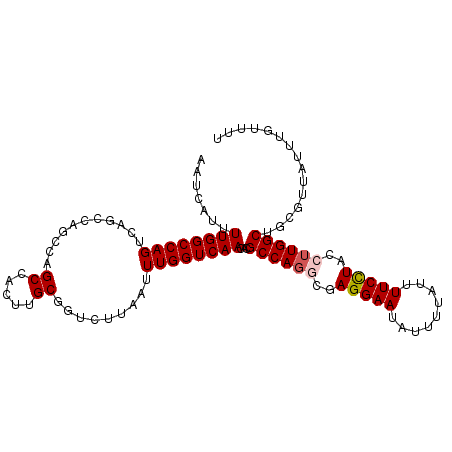
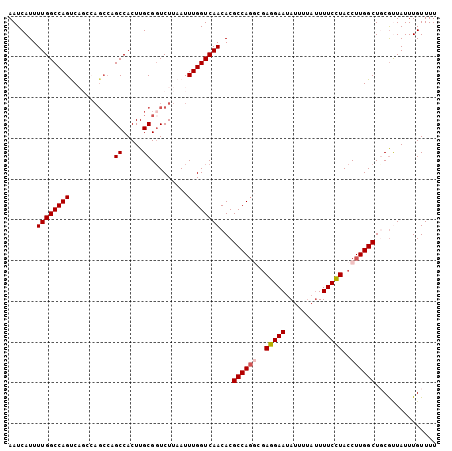
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:41:36 2006