
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 8,532,067 – 8,532,175 |
| Length | 108 |
| Max. P | 0.623321 |

| Location | 8,532,067 – 8,532,175 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 81.02 |
| Mean single sequence MFE | -24.62 |
| Consensus MFE | -11.54 |
| Energy contribution | -12.22 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.79 |
| Structure conservation index | 0.47 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.595550 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
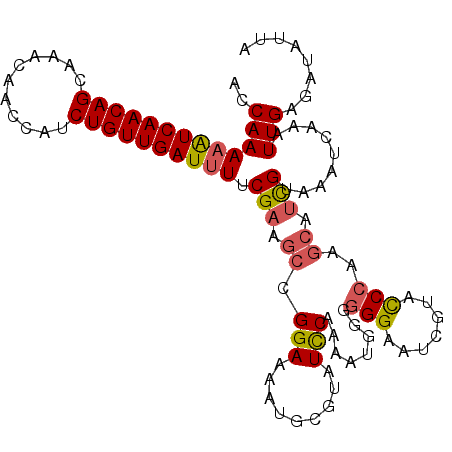
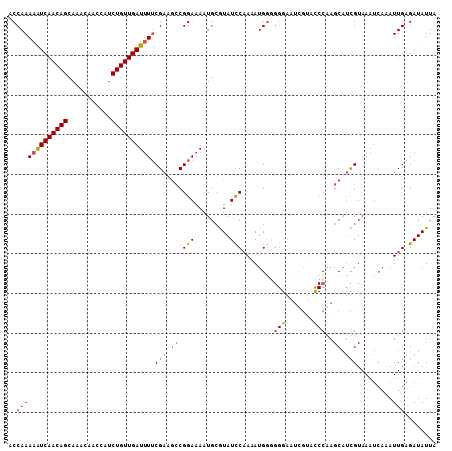
>2L_DroMel_CAF1 8532067 108 + 22407834 UCCAAAAAUCAACAGCAAACAACCAUCUGUUGAUUUUCGAAGCCGGAAAAUGCGUAUCCAAAAUGAGGGGAAUUGAACCCAAGCAUCGUAAAUCAGAUUGAGAUAUUA (((.(((((((((((...........))))))))))).(....))))......((((((((..((((((........)))..((...))...)))..))).))))).. ( -20.90) >DroSec_CAF1 41300 108 + 1 ACCAAAAAUCAACAGCAAACAACCAUCUGUUGAUUUACGAAGCCGGAAAAUGCGUAUUCAAAAUGGGGGGAAUCGUAUCCGAGCAUCGUAAAUCAAAUUGAGAUAUUA ........................((((.(((((((((((.((((((...((((.((((..........)))))))))))).)).)))))))))))....)))).... ( -28.60) >DroSim_CAF1 40615 108 + 1 ACCAAAAAUCAACAGCAAACAACCAUCUGUUGAUUUUCGAAGCCGGAAAAUGAGUAUCCAAAAUGGUGGGAAUCGUACCCGAGCAUCGUAAAUCAAAUUGAGAUAUUA ....(((((((((((...........)))))))))))(((.(((((...((((...((((......))))..))))..))).)).))).................... ( -23.80) >DroEre_CAF1 41192 90 + 1 UCCAAAAGUCAACAGCAAACAAUCAGCUGUUGAUCUUCGGAAACGGAAAAUGCAUAUCCUAUAUGG-------------CAAGCCAUGUAA-----AUUGAGAUAUUA .......(((((((((.........))))))))).((((....))))......(((((((((((((-------------....))))))).-----...).))))).. ( -25.20) >consensus ACCAAAAAUCAACAGCAAACAACCAUCUGUUGAUUUUCGAAGCCGGAAAAUGCGUAUCCAAAAUGGGGGGAAUCGUACCCAAGCAUCGUAAAUCAAAUUGAGAUAUUA ..(((((((((((((...........)))))))))).(((.((.(((.........)))........(((.......)))..)).))).........)))........ (-11.54 = -12.22 + 0.69)

| Location | 8,532,067 – 8,532,175 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 81.02 |
| Mean single sequence MFE | -24.55 |
| Consensus MFE | -15.51 |
| Energy contribution | -15.95 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.623321 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
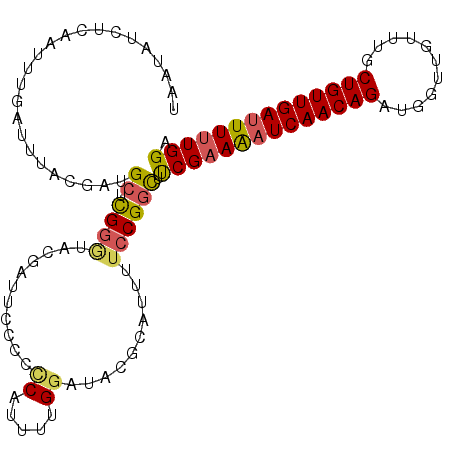
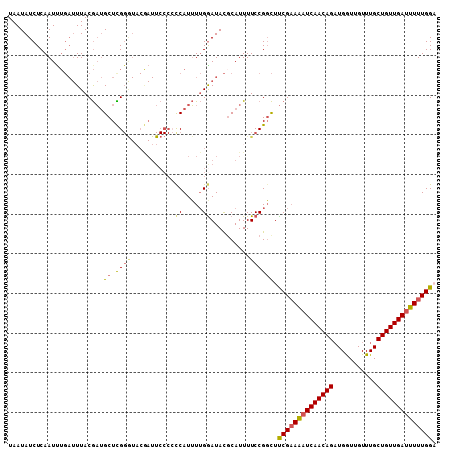
>2L_DroMel_CAF1 8532067 108 - 22407834 UAAUAUCUCAAUCUGAUUUACGAUGCUUGGGUUCAAUUCCCCUCAUUUUGGAUACGCAUUUUCCGGCUUCGAAAAUCAACAGAUGGUUGUUUGCUGUUGAUUUUUGGA ...............(((((.((((...(((........))).)))).)))))..((........))(((((((((((((((...........))))))))))))))) ( -23.40) >DroSec_CAF1 41300 108 - 1 UAAUAUCUCAAUUUGAUUUACGAUGCUCGGAUACGAUUCCCCCCAUUUUGAAUACGCAUUUUCCGGCUUCGUAAAUCAACAGAUGGUUGUUUGCUGUUGAUUUUUGGU .......((((((((((((((((.((.((((..((((((..........)))).)).....)))))).)))))))))))(((((....)))))..)))))........ ( -26.30) >DroSim_CAF1 40615 108 - 1 UAAUAUCUCAAUUUGAUUUACGAUGCUCGGGUACGAUUCCCACCAUUUUGGAUACUCAUUUUCCGGCUUCGAAAAUCAACAGAUGGUUGUUUGCUGUUGAUUUUUGGU ...((((.(((..((......(.....)(((.......)))..))..)))))))..............((((((((((((((...........)))))))))))))). ( -23.40) >DroEre_CAF1 41192 90 - 1 UAAUAUCUCAAU-----UUACAUGGCUUG-------------CCAUAUAGGAUAUGCAUUUUCCGUUUCCGAAGAUCAACAGCUGAUUGUUUGCUGUUGACUUUUGGA ..((((((....-----.((.((((....-------------)))).))))))))............(((((((.((((((((.........)))))))).))))))) ( -25.10) >consensus UAAUAUCUCAAUUUGAUUUACGAUGCUCGGGUACGAUUCCCCCCAUUUUGGAUACGCAUUUUCCGGCUUCGAAAAUCAACAGAUGGUUGUUUGCUGUUGAUUUUUGGA ........................((.((((...........((.....))..........)))))).((((((((((((((...........)))))))))))))). (-15.51 = -15.95 + 0.44)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:41:28 2006