
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 8,501,037 – 8,501,156 |
| Length | 119 |
| Max. P | 0.999649 |

| Location | 8,501,037 – 8,501,156 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.88 |
| Mean single sequence MFE | -25.25 |
| Consensus MFE | -22.78 |
| Energy contribution | -22.82 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.90 |
| SVM decision value | 3.05 |
| SVM RNA-class probability | 0.998278 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
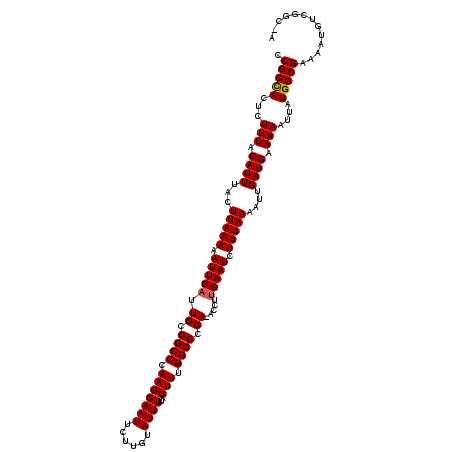
>2L_DroMel_CAF1 8501037 119 + 22407834 CCGGUACUCUUUACAUUUACUUAAAAAUUCAUUGCGGCCCAAGGAAUUCUUGUAUUUUACUUUGGCCCCAAACCUUGAAUCUUUAAAAUUGAUGAAAAAUUAUGCCGAAAAUGUCGGC-A .............((((...(((((.((((((((.((((.(((((((......))))..))).)))).)))....))))).)))))....))))........((((((.....)))))-) ( -24.80) >DroSec_CAF1 10487 119 + 1 CCGGCACUCUUUACAUUUACUUAAAAAUUCAUUGCGGCCCAAGGAAUUCUUGUAUUUUACUUUGGCCCCA-ACCUUGAAUCUUUAAAAUUGAUGAAAAAUUAUGCCGAAAAUAAUGCCGA .((((........((((...(((((.((((((((.((((.(((((((......))))..))).)))).))-)...))))).)))))....))))....(((((.......))))))))). ( -25.10) >DroSim_CAF1 10040 118 + 1 CCGGCACUCUUUACAUUUACUUAAAAAUUCCUUGCGGCCCAAGGAAUUCUUGUAUUUUACUUUGGCCCCA-ACCUUGAAUCUUUAAAAUUGAUGAAAAAUUAUGCCGAAAAUGUCGGC-A ((((((...............(((.(((((((((.....))))))))).)))........((((((....-.......(((.........)))..........))))))..)))))).-. ( -23.60) >DroEre_CAF1 10069 119 + 1 CCGGCACUCUUUGCAUUUACUUAAAAAUUCAUUGCGGCCCAAGGAAUUCUUUUAUUUCACUUCGGCCCCAAACCUUGAAUCUUUAAAAUUGAUGAAAAAUUAUGCCGAAAAUGUCGGC-A .(((((.......((((...(((((.((((((((.((((.((((((.........))).))).)))).)))....))))).)))))....))))........)))))...........-. ( -26.76) >DroYak_CAF1 10670 118 + 1 UCGGCACUCUUUACAUUUACUUAAAAAUUCAUUGCGGCCCAAGGAAUUCUUUUAUUUUACUUCGGCCCCA-ACCUUGAAUCUUUAAAAUUGAUGAAAAAUUAUGCCGAAAAUGCCGGC-A ((((((...(((.((((...(((((.((((((((.((((.(((((((......))))..))).)))).))-)...))))).)))))....)))).)))....))))))..........-. ( -26.00) >consensus CCGGCACUCUUUACAUUUACUUAAAAAUUCAUUGCGGCCCAAGGAAUUCUUGUAUUUUACUUUGGCCCCA_ACCUUGAAUCUUUAAAAUUGAUGAAAAAUUAUGCCGAAAAUGUCGGC_A .(((((...(((.((((...(((((.(((((.((.((((.(((((((......))))..))).)))).)).....))))).)))))....)))).)))....)))))............. (-22.78 = -22.82 + 0.04)


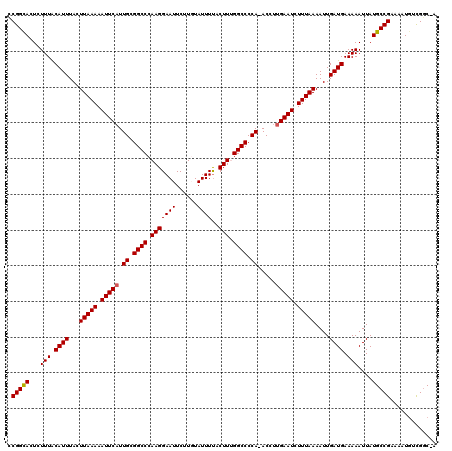
| Location | 8,501,037 – 8,501,156 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.88 |
| Mean single sequence MFE | -32.98 |
| Consensus MFE | -29.03 |
| Energy contribution | -29.27 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.83 |
| Structure conservation index | 0.88 |
| SVM decision value | 3.83 |
| SVM RNA-class probability | 0.999649 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
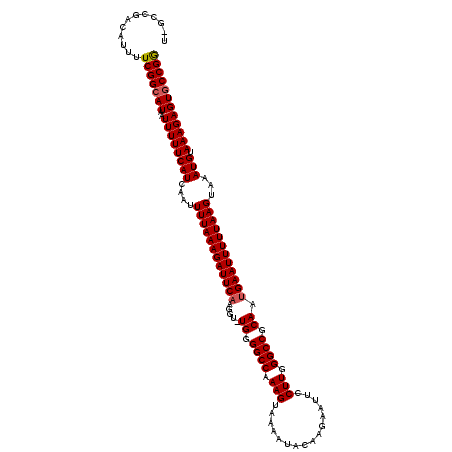
>2L_DroMel_CAF1 8501037 119 - 22407834 U-GCCGACAUUUUCGGCAUAAUUUUUCAUCAAUUUUAAAGAUUCAAGGUUUGGGGCCAAAGUAAAAUACAAGAAUUCCUUGGGCCGCAAUGAAUUUUUAAGUAAAUGUAAAGAGUACCGG (-(((((.....))))))...((((((((....((((((((((((....(((.((((..(......).((((.....)))))))).)))))))))))))))...))).)))))....... ( -30.20) >DroSec_CAF1 10487 119 - 1 UCGGCAUUAUUUUCGGCAUAAUUUUUCAUCAAUUUUAAAGAUUCAAGGU-UGGGGCCAAAGUAAAAUACAAGAAUUCCUUGGGCCGCAAUGAAUUUUUAAGUAAAUGUAAAGAGUGCCGG .............((((((..((((((((....((((((((((((...(-((.((((..(......).((((.....)))))))).)))))))))))))))...))).))))))))))). ( -31.90) >DroSim_CAF1 10040 118 - 1 U-GCCGACAUUUUCGGCAUAAUUUUUCAUCAAUUUUAAAGAUUCAAGGU-UGGGGCCAAAGUAAAAUACAAGAAUUCCUUGGGCCGCAAGGAAUUUUUAAGUAAAUGUAAAGAGUGCCGG .-...........((((((..((((((((....(((((((((((....(-((.((((..(......).((((.....)))))))).))).)))))))))))...))).))))))))))). ( -31.10) >DroEre_CAF1 10069 119 - 1 U-GCCGACAUUUUCGGCAUAAUUUUUCAUCAAUUUUAAAGAUUCAAGGUUUGGGGCCGAAGUGAAAUAAAAGAAUUCCUUGGGCCGCAAUGAAUUUUUAAGUAAAUGCAAAGAGUGCCGG .-...........((((((..((((((((....((((((((((((....(((.((((.(((.(((.........)))))).)))).)))))))))))))))...))).))))))))))). ( -36.10) >DroYak_CAF1 10670 118 - 1 U-GCCGGCAUUUUCGGCAUAAUUUUUCAUCAAUUUUAAAGAUUCAAGGU-UGGGGCCGAAGUAAAAUAAAAGAAUUCCUUGGGCCGCAAUGAAUUUUUAAGUAAAUGUAAAGAGUGCCGA .-..(((((((((..((((..............((((((((((((...(-((.((((.(((................))).)))).)))))))))))))))...))))..))))))))). ( -35.62) >consensus U_GCCGACAUUUUCGGCAUAAUUUUUCAUCAAUUUUAAAGAUUCAAGGU_UGGGGCCAAAGUAAAAUACAAGAAUUCCUUGGGCCGCAAUGAAUUUUUAAGUAAAUGUAAAGAGUGCCGG ............(((((((..((((((((....((((((((((((.....((.((((.(((................))).)))).)).))))))))))))...))).)))))))))))) (-29.03 = -29.27 + 0.24)

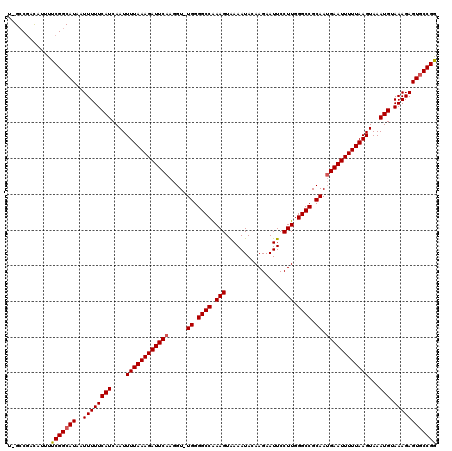
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:40:44 2006