
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 8,495,966 – 8,496,126 |
| Length | 160 |
| Max. P | 0.969594 |

| Location | 8,495,966 – 8,496,086 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.81 |
| Mean single sequence MFE | -38.67 |
| Consensus MFE | -34.72 |
| Energy contribution | -34.85 |
| Covariance contribution | 0.13 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.42 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.628069 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
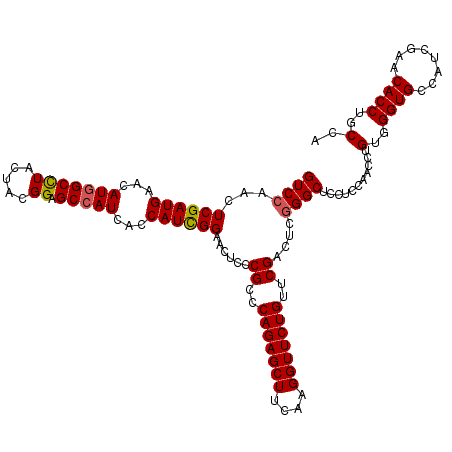
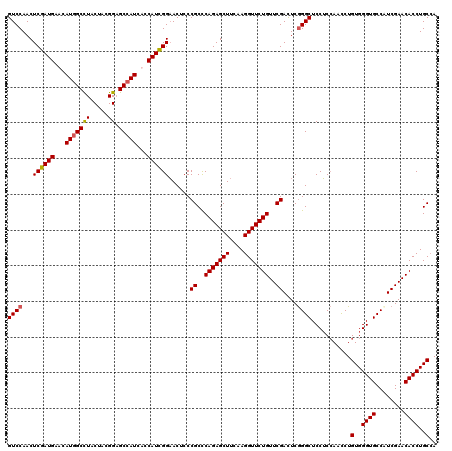
>2L_DroMel_CAF1 8495966 120 - 22407834 GUCCAACUCGAUGAAUAUGGCUUACUACGGAGCCAUCUCCAUCGGAACUCCCGCUCAGAGCUUCAAGGUUCUGUUCGACUCAGGCUCCUCGAACCUGUGGGUGCCAUCGAACACCUGCAA (((....((((((...(((((((......)))))))...))))))......((..(((((((....)))))))..)).....)))..........((..((((........))))..)). ( -37.60) >DroSec_CAF1 5116 120 - 1 GUCCAACUCGAUGAACAUGGCCUACUACGGAGCCAUCUCCAUUGGAACUCCCGCCCAGAGCUUCAAGGUUCUGUUCGACUCGGGCUCCUCCAAUCUGUGGGUGCCUUCGAACACCUGCCA ........(((((...(((((((.....)).)))))...)))))((((((.......))).)))..(((..(((((((...((((.((..(.....)..)).)))))))))))...))). ( -36.90) >DroEre_CAF1 4981 120 - 1 GUCCAACUCGAUGAACAUGGCCUACUACGGAGCCAUCAGCAUCGGAACGCCCGCCCAGAGCUUCAAGGUUCUGUUCGAUUCGGGCUCCUCCAACCUGUGGGUGCCCUCGAACACCUGCCA ........(((((...(((((((.....)).)))))...)))))(((.((.(.....).)))))..(((..(((((((...((((.((..(.....)..)).)))))))))))...))). ( -41.80) >DroYak_CAF1 5360 120 - 1 GUCCAACUCGAUGAACAUGGCCUACUAUGGUGCGAUCAGCAUCGGCACUCCCGCCCAGAGCUUCAAGGUUCUGUUCGACUCGGGCUCCGCCAAUCUGUGGGUGCCAUCGAACACCUGCCA .......((((((((((..((((.....(((((.....)))))(((......)))..........))))..)))))......((((((((......))))).)))))))).......... ( -38.40) >consensus GUCCAACUCGAUGAACAUGGCCUACUACGGAGCCAUCACCAUCGGAACUCCCGCCCAGAGCUUCAAGGUUCUGUUCGACUCGGGCUCCUCCAACCUGUGGGUGCCAUCGAACACCUGCCA ((((...((((((...(((((((.....)).)))))...))))))......((..(((((((....)))))))..))....))))...........(..((((........))))..).. (-34.72 = -34.85 + 0.13)

| Location | 8,496,006 – 8,496,126 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.36 |
| Mean single sequence MFE | -45.27 |
| Consensus MFE | -40.44 |
| Energy contribution | -41.75 |
| Covariance contribution | 1.31 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.64 |
| SVM RNA-class probability | 0.969594 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
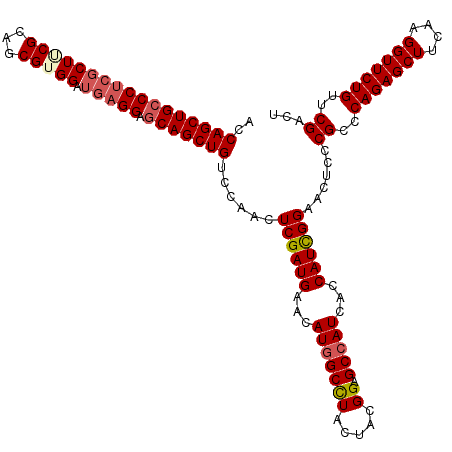
>2L_DroMel_CAF1 8496006 120 - 22407834 ACCAGCUGCCCUCGCUUCGCAGCGUGGAUGAGGAACAGCUGUCCAACUCGAUGAAUAUGGCUUACUACGGAGCCAUCUCCAUCGGAACUCCCGCUCAGAGCUUCAAGGUUCUGUUCGACU ..((((((.(((((.(((((...))))))))))..))))))......((((((...(((((((......)))))))...))))))......((..(((((((....)))))))..))... ( -44.20) >DroSec_CAF1 5156 120 - 1 ACCAGCUGCCCUCGCUUCGCAGCGUGGAUGAGGAGCAGCUGUCCAACUCGAUGAACAUGGCCUACUACGGAGCCAUCUCCAUUGGAACUCCCGCCCAGAGCUUCAAGGUUCUGUUCGACU ..((((((((((((.(((((...)))))))))).)))))))......((((((...(((((((.....)).)))))...))))))......((..(((((((....)))))))..))... ( -45.30) >DroEre_CAF1 5021 120 - 1 ACCAGCUGCCCUCGCUACGCAGCGUGGAUGAGGAGCAGCUGUCCAACUCGAUGAACAUGGCCUACUACGGAGCCAUCAGCAUCGGAACGCCCGCCCAGAGCUUCAAGGUUCUGUUCGAUU ..(((((((((((((((((...))))).))))).)))))))......((((((...(((((((.....)).)))))...))))))......((..(((((((....)))))))..))... ( -50.60) >DroYak_CAF1 5400 120 - 1 ACCAGCUGCCCAGCCUGCGCAGCGUGGAUGAGGAGCAGCUGUCCAACUCGAUGAACAUGGCCUACUAUGGUGCGAUCAGCAUCGGCACUCCCGCCCAGAGCUUCAAGGUUCUGUUCGACU ..(((((((((..(((((...))).))....)).)))))))..........((((((..((((.....(((((.....)))))(((......)))..........))))..))))))... ( -41.00) >consensus ACCAGCUGCCCUCGCUUCGCAGCGUGGAUGAGGAGCAGCUGUCCAACUCGAUGAACAUGGCCUACUACGGAGCCAUCACCAUCGGAACUCCCGCCCAGAGCUUCAAGGUUCUGUUCGACU ..(((((((((((((((((...))))).))))).)))))))......((((((...(((((((.....)).)))))...))))))......((..(((((((....)))))))..))... (-40.44 = -41.75 + 1.31)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:40:31 2006