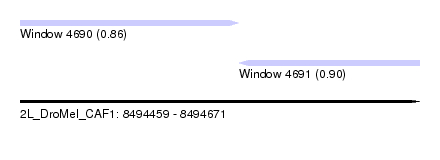
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 8,494,459 – 8,494,671 |
| Length | 212 |
| Max. P | 0.898193 |

| Location | 8,494,459 – 8,494,575 |
|---|---|
| Length | 116 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.01 |
| Mean single sequence MFE | -33.00 |
| Consensus MFE | -25.75 |
| Energy contribution | -26.03 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.81 |
| SVM RNA-class probability | 0.856649 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 8494459 116 + 22407834 ----CGGAUGCCCCUGCAUCCUUCGGCCGCUACUCCUAAAUGGGUGUUGUCACUGUUGUGUUUGCAGUUGCAAAACUAUAGACCCGCACAAUUCUCUGACAUCACUGGCUAUCCUUUGAC ----.((((((....))))))...((((......((.....))(((.(((((..(((((((((((....))))............)))))))....))))).))).)))).......... ( -33.80) >DroSec_CAF1 3639 114 + 1 ----CGGAUGCCGC-GCGUCCUUCGGCCGCUACUCCUAAAUG-GUGUUGUCACUGUUGUGUUUGCAGUUGCAAAACUAUAGUCCCGCACAAUUCUCUGACAUCACUGGCUAUCCUUUGGC ----.(((((((((-(.(((....))))))...........(-(((.(((((..(((((((((((....)))).((....))...)))))))....))))).))))))).))))...... ( -35.20) >DroSim_CAF1 3653 114 + 1 ----CGGAUGCCGC-GCGUCCUUCAGCCGCUACUCCUAAAUG-GUGUUGUCACUGUUGUGUUUGCAGUUGCAAAACUAUAGACCCGCACAAUUCUCUGACAUCACUGGCUAUCCUUUGAC ----.(((((((((-(.((......)))))...........(-(((.(((((..(((((((((((....))))............)))))))....))))).))))))).))))...... ( -33.10) >DroEre_CAF1 3260 118 + 1 CCAUCGGAUGCCCC-UCGUCCUUCUGCCGCUGCUCCCAAAUG-GUGUUGUCACUGUUGUGUUUGCUGUCGCAAAACUAUAGUCCCGCACAAUUCUCUGACAUCAGUGGCUAUCCUUUGAC .....(((((....-.)))))....(((((((...((....)-)...(((((..(((((((..(((((.(.....).)))))...)))))))....))))).)))))))........... ( -31.50) >DroYak_CAF1 3651 114 + 1 ----CGGAUGCCCC-UCGUCCUUCUGCCGCUACUCCCAAAUG-GUGUUGUCACUAUUGUGUUUGCUGUGGCAAAACUAUAGUCCCGCACAAUUCCCUGACAUCACUGGCUAUCCUUUGAC ----.(((((((..-..((......))..............(-(((.(((((..(((((((..(((((((.....)))))))...)))))))....))))).))))))).))))...... ( -31.40) >consensus ____CGGAUGCCCC_GCGUCCUUCGGCCGCUACUCCUAAAUG_GUGUUGUCACUGUUGUGUUUGCAGUUGCAAAACUAUAGUCCCGCACAAUUCUCUGACAUCACUGGCUAUCCUUUGAC .....((((((....))))))....((((..............(((.(((((..(((((((((((....))))............)))))))....))))).)))))))........... (-25.75 = -26.03 + 0.28)



| Location | 8,494,575 – 8,494,671 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 72.75 |
| Mean single sequence MFE | -39.66 |
| Consensus MFE | -25.73 |
| Energy contribution | -26.13 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.65 |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.898193 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 8494575 96 - 22407834 GCCUGUUGGCCGGGCAAUGAUGUGUGUCACCGGGUGUCAAGUGGG------------------------GACUCUGGUAGCCCGGGGGAAUGCCACAGAUGUCAGGUAUACUGCCGGGGU (((((.....)))))...(((....)))((((((.(((.......------------------------)))))))))..(((((.((.(((((((....))..))))).)).))))).. ( -36.10) >DroSec_CAF1 3753 83 - 1 GCCCGUUGCCCGGGCAAUGAUGUGUGUCACCGGGUGUCAAGAGGG------------------------GACUUUGCUAGCCCGGGGGAAUGCCACAAAUGUCAGG-------------U (((((.....)))))..((((((.(((..((((((....((((..------------------------..))))....))))))((.....))))).))))))..-------------. ( -34.70) >DroSim_CAF1 3767 83 - 1 GCCUGUUGCCCGGGCAAUGAUGUGUGUCACCGGGUGUCAAGAGGG------------------------GACUUUGCUAGCCCGGGGGAAUGCCACAGAUGUCAGG-------------U (((((((((....)))))(((((.(((..((((((....((((..------------------------..))))....))))))((.....))))).))))))))-------------) ( -35.10) >DroEre_CAF1 3378 114 - 1 GCCUGUUGCCCGGGCAAUGAAGUGUGUCACCGGGUGUCAGGAGUGUCAGUAGUGUCAGGAGGC------GACUGUGGAACCUCGGGGGAAUGCCACAGAUGUCGGGGAUAGUGCUGGGGU .((((.((((((((((........)))..))))))).))))..(.((((((.((((....(((------(.((((((..((....)).....)))))).))))...)))).)))))).). ( -46.90) >DroYak_CAF1 3765 114 - 1 GCCUGUUGCCCGGGCAAUGAAGUGUGUCACCGAGUGUCAGGAGGGA------ACCCUGGAGGCUCUGAGAACUCGGAGACCUCGGGGGAAUGCCACAGAUGUCAGGGAUAGUGGCGGGGU .(((((..(...((((........)))).((..(..((.(...((.------.((((.((((((((((....)))))).)))).))))....)).).))..)..))....)..))))).. ( -45.50) >consensus GCCUGUUGCCCGGGCAAUGAUGUGUGUCACCGGGUGUCAAGAGGG________________________GACUCUGCUAGCCCGGGGGAAUGCCACAGAUGUCAGG_AUA_UG__GGGGU (((((.....)))))..((((((.(((..((((((....((((............................))))....))))))((.....))))).))))))................ (-25.73 = -26.13 + 0.40)


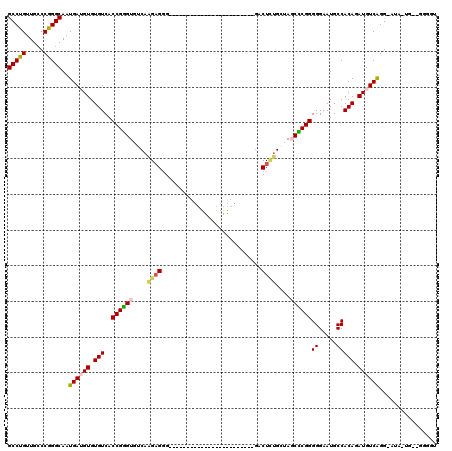
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:40:28 2006