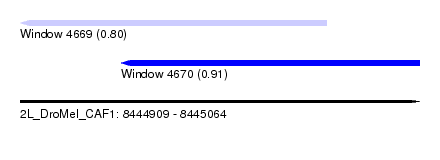
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 8,444,909 – 8,445,064 |
| Length | 155 |
| Max. P | 0.914732 |

| Location | 8,444,909 – 8,445,028 |
|---|---|
| Length | 119 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 79.44 |
| Mean single sequence MFE | -27.35 |
| Consensus MFE | -15.75 |
| Energy contribution | -15.12 |
| Covariance contribution | -0.63 |
| Combinations/Pair | 1.61 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.799754 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 8444909 119 - 22407834 GCUGAGGAUCAGAUUUAUAUUUCUUAUAGAUUUCCAUUUCUUCCGCACCCAUGGGUACCUGUCUACCCAUGAUCAUGCGUUCCAUUUCCUCCACAUCCUUGUCAAAGGCCCUUGACCUC ...(((((..(((((((((.....)))))))))..........((((..((((((((......))))))))....)))).......))))).........(((((......)))))... ( -29.50) >DroSec_CAF1 4576 119 - 1 GCUGAGGAUCAGAUUUAUAUUUCUUAUAUAUUUCCAUUUCUUCCGCACUCAUGGGUACCUGUCUACCCAUGAUCAUGCGCUCCAUUUCCUCCACAUUCUUGUCAAAGGCCCUUGACCUC ...(((((...((..(((((.....)))))..)).........((((.(((((((((......)))))))))...)))).......))))).........(((((......)))))... ( -30.10) >DroSim_CAF1 4131 119 - 1 GCUGAGGAUCAGAUUUAUAUUUCUUAUAGAUUUCCAUUUCUUCCGCACUCAUGGGUACCUGUCUACCCAUGAUCAUGCGCUCCAUUUCCUCCACAUUCUUGUCAAAGGCCCUUGACCUC ...(((((..(((((((((.....)))))))))..........((((.(((((((((......)))))))))...)))).......))))).........(((((......)))))... ( -31.30) >DroEre_CAF1 4236 119 - 1 GCUGAGGGUCGCAUGUAUACUUCCUAAAGAUUUCAACUUCUUCCUCGCUUAUGGGUACCUGUCUAUUCAUGAUCAUGAGCUCCAUUUGCCCCACAUCCUUGUCGAAAGCCCUAGACCUC .....(((((................((((........))))....((((..(((((..((....(((((....)))))...))..))))).(((....)))...))))....))))). ( -20.30) >DroYak_CAF1 3985 119 - 1 GCUGAGGGUCAGAUUUAUAUUUCUUGAAGAUUUCCACUUCCUCCGCACUUGUGGGUACCUGUCUACCCAUGAUCAUGUGCUCCAUUUGCUCCACAUCUUUGUCAAAAGCACCAGACCUC .....(((((...............((((.......))))....(((((..((((((......))))))..)....))))......((((..(((....)))....))))...))))). ( -26.00) >DroAna_CAF1 3486 119 - 1 GGGUGGGAUCUGAAUCGUAUUUCUUAAAAAGCUGGAUCUCUUCAUCACUUUCGGGCACCUGCCUGCUCAUGAUCAUCUGGCCCAUGCUCUCCAGUUGUUUGUCCGAGAUACUCGAGUUU .....(((..(((...(((((((.((((.(((((((.......((((....(((((....)))))....)))).....(((....))).))))))).))))...))))))))))..))) ( -26.90) >consensus GCUGAGGAUCAGAUUUAUAUUUCUUAAAGAUUUCCAUUUCUUCCGCACUCAUGGGUACCUGUCUACCCAUGAUCAUGCGCUCCAUUUCCUCCACAUCCUUGUCAAAGGCCCUUGACCUC ...((((.((.................................((((.(((((((((......)))))))))...)))).................((((....)))).....)))))) (-15.75 = -15.12 + -0.63)



| Location | 8,444,948 – 8,445,064 |
|---|---|
| Length | 116 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 77.13 |
| Mean single sequence MFE | -31.36 |
| Consensus MFE | -21.98 |
| Energy contribution | -20.10 |
| Covariance contribution | -1.88 |
| Combinations/Pair | 1.68 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.70 |
| SVM decision value | 1.10 |
| SVM RNA-class probability | 0.914732 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 8444948 116 - 22407834 UGGGUUUGCCGGACGUUAGUAUCAGUAUGUUGUAGUGCUGAGGAUCAGAUUUAUAUUUCUUAUAGAUUUCCAUUUCUUCCGCACCCAUGGGUACCUGUCUACCCAUGAUCAUGCGU .((((..((.(((........(((((((......)))))))(((...((((((((.....)))))))))))......)))))))))(((((((......))))))).......... ( -33.50) >DroSec_CAF1 4615 116 - 1 UGGGUUUACCCGACGUUAGUAUCAGUAUGUUGUAGUGCUGAGGAUCAGAUUUAUAUUUCUUAUAUAUUUCCAUUUCUUCCGCACUCAUGGGUACCUGUCUACCCAUGAUCAUGCGC .(((....))).............(((((.....((((.(((((...((..(((((.....)))))..))....))))).))))(((((((((......))))))))).))))).. ( -37.40) >DroSim_CAF1 4170 116 - 1 UGGGUUUGCCGGACGUUAGUAUCAGUAUGUUGUAGUGCUGAGGAUCAGAUUUAUAUUUCUUAUAGAUUUCCAUUUCUUCCGCACUCAUGGGUACCUGUCUACCCAUGAUCAUGCGC ..((....))(((........(((((((......)))))))(((...((((((((.....)))))))))))......)))(((.(((((((((......)))))))))...))).. ( -35.30) >DroEre_CAF1 4275 116 - 1 UGGGUUUGCCGAACGUUAGUAUCAGUACGUUGUAGUGCUGAGGGUCGCAUGUAUACUUCCUAAAGAUUUCAACUUCUUCCUCGCUUAUGGGUACCUGUCUAUUCAUGAUCAUGAGC ..((....))((.(((.(((((((((((......)))))))(((((.((((...........((((........)))).......)))).).))))...)))).))).))...... ( -22.27) >DroWil_CAF1 5997 113 - 1 UUGGUUCUAUG---GCCAUGAUCAGUAUAUUAUAAUGCUGAGGAUCAAGUGCGUAUUCAUUAAGCAAAUCACGAUCCUCCUGUCUGUUGGGCCAUUCCCUGUUCAUUAUCAACUUG ..((....(((---(((....(((((((......)))))))(((((...(((.((.....)).)))......)))))............)))))).)).................. ( -27.00) >DroYak_CAF1 4024 116 - 1 UGGGUUUGCCGACAUUAAGUAUCAGCACGUUGUAGUGCUGAGGGUCAGAUUUAUAUUUCUUGAAGAUUUCCACUUCCUCCGCACUUGUGGGUACCUGUCUACCCAUGAUCAUGUGC ((((((((((.((.....)).(((((((......))))))).)).))))))))........((((.......))))....(((((..((((((......))))))..)....)))) ( -32.70) >consensus UGGGUUUGCCGGACGUUAGUAUCAGUAUGUUGUAGUGCUGAGGAUCAGAUUUAUAUUUCUUAAAGAUUUCCAUUUCUUCCGCACUCAUGGGUACCUGUCUACCCAUGAUCAUGCGC ..((....))...........(((((((......)))))))......................................((((.(((((((((......)))))))))...)))). (-21.98 = -20.10 + -1.88)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:40:08 2006