
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 8,435,258 – 8,435,378 |
| Length | 120 |
| Max. P | 0.582556 |

| Location | 8,435,258 – 8,435,378 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 76.67 |
| Mean single sequence MFE | -38.35 |
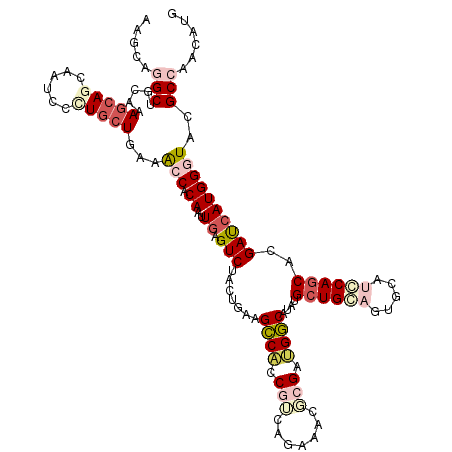
| Consensus MFE | -20.58 |
| Energy contribution | -22.42 |
| Covariance contribution | 1.84 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.54 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.582556 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
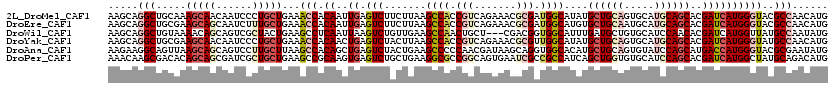
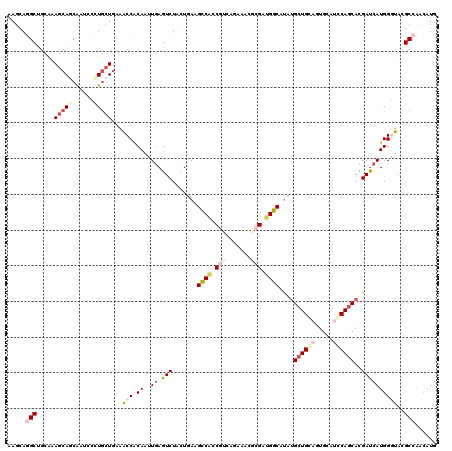
>2L_DroMel_CAF1 8435258 120 - 22407834 AAGCAGGCUGCAAAGCAACAAUCCCUGCUGAAACCACAAUUGAGUCUUCUUAAGCCACCGUCAGAAACGCGAUGGCAUAUGCUGCAGUGCAUGCAGCACGAUCAUGGGUACGCCAACAUG .....((((((..((((........))))....(((..((((...........((((.(((.......))).))))...(((((((.....)))))))))))..)))))).)))...... ( -36.10) >DroEre_CAF1 8220 120 - 1 AAGCAGGCUGCGAAGCAGCAAUCUUUGCUGAAACCACAAUUGAGUCUUCUUAAGCCACCGUCAGAAACGCGAUGGCAUGUGCUGCAAUGCAUGCAGCACGAUCAUGGGUACGCCAACAUG .....((((((....(((((.....)))))...(((...(((((....)))))((((.(((.......))).)))).(((((((((.....)))))))))....)))))).)))...... ( -40.30) >DroWil_CAF1 9568 117 - 1 AAGCAGGCUGUAAAACAGCAGUCGCUACUGAAGCCUCAAUUAAGUCUGUUGAAGCCAACUGCU---CGACGGUGGCAUUUGAUGCUGUGCAUCCAACACGAUCAUGGUUAUGCCAAUAUG .(((.((((((......)))))))))...................(((((((.((.....)))---))))))((((((.....((((((.(((......))))))))).))))))..... ( -32.70) >DroYak_CAF1 8322 120 - 1 AAGCAGGCUGCGAAGCAACAAUCCCUGCUGAAACCACAACUGAGUCUACUUAAGCCACCGUCAGAAACGCGUUGGCAUAUGCUGCAGUGCAUGCAGCACGAUCAUGGGUAUGCCAACAUG ..((.((((....((((........))))...........((((....))))))))...((.....))))((((((((((.(((..((((.....))))...)).).))))))))))... ( -35.80) >DroAna_CAF1 8669 120 - 1 AAGAAGGCAGUUAAGCAGCAGUCCUUGCUUAAGCCACAGCUGAGUCUACUGAAGCCCCCAACGAUAAGCAGGUGGCACAUGCUGCAGUGUAUCCAGCAUGACCAUGGGUACGCGAAUAUG ......((((((((((((......))))))).(((((.(((..(((...((.......))..))).)))..)))))....))))).(((((((((.........)))))))))....... ( -37.80) >DroPer_CAF1 8039 120 - 1 AAACAAGCGACACAGCAGCGAUCGCUGCUGAAGCCGCAAGUGAGUCUGCUGAAGGCGCCGGCAGUGAAUCGCCGCCAUCAGCUGGUGUGCAUCCAGCACGAUCAUGGCUAUGCAGACAUG ......(((.(.(((((((....)))))))..).)))......((((((.((.((((.(((.......))).)))).))(((((.(((((.....)))))....)))))..))))))... ( -47.40) >consensus AAGCAGGCUGCAAAGCAGCAAUCCCUGCUGAAACCACAAUUGAGUCUACUGAAGCCACCGUCAGAAACGCGAUGGCAUAUGCUGCAGUGCAUCCAGCACGAUCAUGGGUACGCCAACAUG .....(((.....(((((......)))))...(((.((..((.(((.......((((.(((.......))).))))....((((((.....))))))..))))))))))..)))...... (-20.58 = -22.42 + 1.84)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:40:03 2006