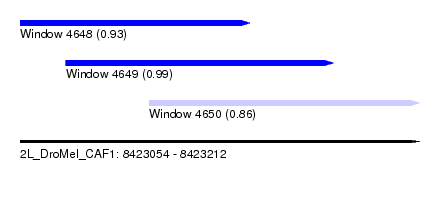

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 8,423,054 – 8,423,212 |
| Length | 158 |
| Max. P | 0.990683 |

| Location | 8,423,054 – 8,423,145 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 95.66 |
| Mean single sequence MFE | -29.65 |
| Consensus MFE | -25.39 |
| Energy contribution | -25.67 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.930340 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
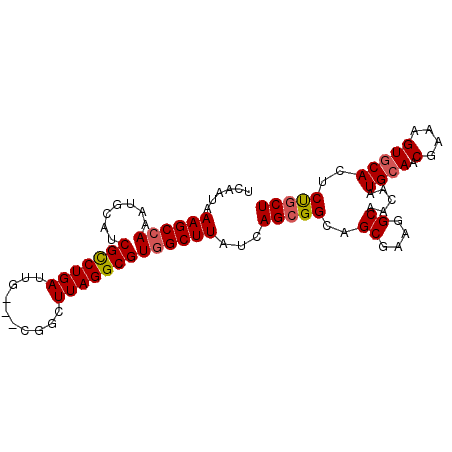
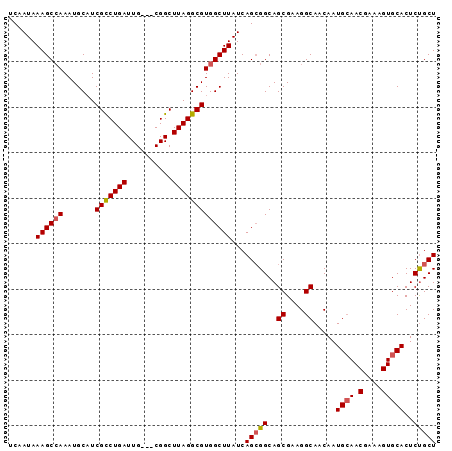
>2L_DroMel_CAF1 8423054 91 + 22407834 UCAAUAAAGCCAAAUGCAUCGUCUGAUUG---CGGCUUAGGCGUGGCUUAUCAGCGGCAGCGAAGGCAACAAUGCAACGAAAGUGCACUCUGCU ......((((((.......(((((((...---....)))))))))))))...(((((..((....)).....((((.(....)))))..))))) ( -26.81) >DroSec_CAF1 21246 91 + 1 UCAAUAAAGCCAAAUGCAUCGCCUGAUUG---CGGCUUAGGCGUGGCUUAUCAGCGGCAGCGAAGGCAACAAUGGAACGAAAGUGCACUCCGCU ......((((((.......(((((((...---....)))))))))))))...(((((..((....)).....((..((....)).))..))))) ( -28.41) >DroSim_CAF1 21061 91 + 1 UCAAUAAAGCCAAAUGCAUCGCCUGAUUG---CGGCUUAGGCGUGGCUUAUCAGCGGCAGCGAAGGCAACAAUGCAACGAAAGUGCACUCUGCU ......((((((.......(((((((...---....)))))))))))))...(((((..((....)).....((((.(....)))))..))))) ( -29.51) >DroEre_CAF1 21481 94 + 1 UCAAUAAAGCCAAAUGCUUCGCCUGAUUGCUGCGGCUUAGGCGUGGCUUAUCAGAGGCAGCGAAGGCAGCAAUGCAACGAAAGUGCACUCUGCU ......((((((.......(((((((..((....)))))))))))))))..(((((((.((....)).))..((((.(....)))))))))).. ( -35.21) >DroYak_CAF1 21509 91 + 1 UCAAUAAAGCGAAAUGCAUCGCCUGAUUG---CGGCUUAGGCGUGGCUUAUCAGCGGCAGCGAAGGCAACAAUGCAACGAAAGUGCACUCUGCU ........((((...((..(((((((...---....)))))))..))...)).))(((((.(..(....)..((((.(....)))))).))))) ( -28.30) >consensus UCAAUAAAGCCAAAUGCAUCGCCUGAUUG___CGGCUUAGGCGUGGCUUAUCAGCGGCAGCGAAGGCAACAAUGCAACGAAAGUGCACUCUGCU ......((((((.......(((((((..........)))))))))))))...(((((..((....)).....((((.(....)))))..))))) (-25.39 = -25.67 + 0.28)

| Location | 8,423,072 – 8,423,178 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 94.59 |
| Mean single sequence MFE | -39.76 |
| Consensus MFE | -33.42 |
| Energy contribution | -34.30 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.84 |
| SVM decision value | 2.23 |
| SVM RNA-class probability | 0.990683 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
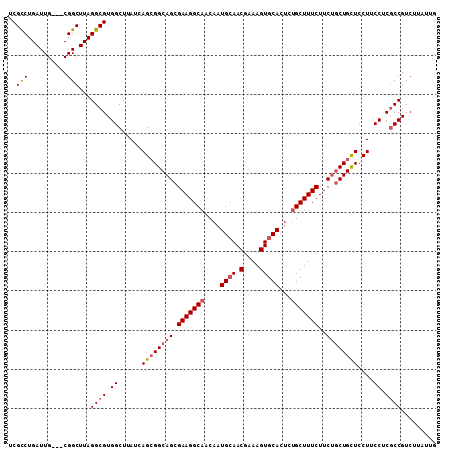
>2L_DroMel_CAF1 8423072 106 + 22407834 UCGUCUGAUUG---CGGCUUAGGCGUGGCUUAUCAGCGGCAGCGAAGGCAACAAUGCAACGAAAGUGCACUCUGCUUUCUUCUACUGCUCCUUCCUCGCCGUCUUAUUG ((((......)---)))....((((.((......(((((.((.(((((((....((((.(....)))))...)))))))..)).)))))....)).))))......... ( -32.20) >DroSec_CAF1 21264 106 + 1 UCGCCUGAUUG---CGGCUUAGGCGUGGCUUAUCAGCGGCAGCGAAGGCAACAAUGGAACGAAAGUGCACUCCGCUUUCUUCUGCUGCUCCUGCCUCGCCGUCUUAUUG ..(((......---.)))...((((.(((.....((((((.((....))......((((.(((((((.....)))))))))))))))))...))).))))......... ( -41.10) >DroSim_CAF1 21079 106 + 1 UCGCCUGAUUG---CGGCUUAGGCGUGGCUUAUCAGCGGCAGCGAAGGCAACAAUGCAACGAAAGUGCACUCUGCUUUCUUCUGCUGCUCCUGCCUCGCCGUCUUAUUG ..(((......---.)))...((((.(((.....((((((((.(((((((....((((.(....)))))...)))))))..))))))))...))).))))......... ( -45.80) >DroEre_CAF1 21499 109 + 1 UCGCCUGAUUGCUGCGGCUUAGGCGUGGCUUAUCAGAGGCAGCGAAGGCAGCAAUGCAACGAAAGUGCACUCUGCUUUCUUCUGCUGUUCCUUCCUCUCCGUCUUAUUG .(((((((..((....))))))))).((......((..((((((((((.((((.((((.(....)))))...)))).))))).)))))..))......))......... ( -38.20) >DroYak_CAF1 21527 106 + 1 UCGCCUGAUUG---CGGCUUAGGCGUGGCUUAUCAGCGGCAGCGAAGGCAACAAUGCAACGAAAGUGCACUCUGCUUUCUUCGGCUGCUCCUUCCGCGCCGUCUUAUUG ..(((......---.)))...(((((((......((((((...(((((((....((((.(....)))))...)))))))....))))))....)))))))......... ( -41.50) >consensus UCGCCUGAUUG___CGGCUUAGGCGUGGCUUAUCAGCGGCAGCGAAGGCAACAAUGCAACGAAAGUGCACUCUGCUUUCUUCUGCUGCUCCUUCCUCGCCGUCUUAUUG ..(((..........)))...((((.((......((((((((.(((((((....((((.(....)))))...)))))))..))))))))....)).))))......... (-33.42 = -34.30 + 0.88)
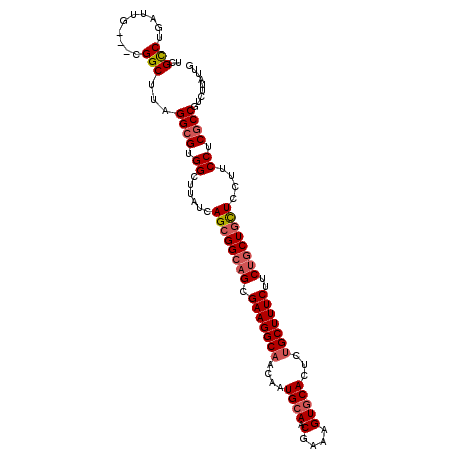


| Location | 8,423,105 – 8,423,212 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 95.33 |
| Mean single sequence MFE | -30.02 |
| Consensus MFE | -25.88 |
| Energy contribution | -26.32 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.84 |
| SVM RNA-class probability | 0.864718 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
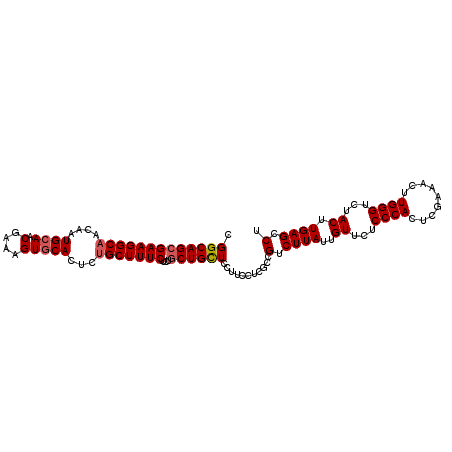
>2L_DroMel_CAF1 8423105 107 + 22407834 CGGCAGCGAAGGCAACAAUGCAACGAAAGUGCACUCUGCUUUCUUCUACUGCUCCUUCCUCGCCGUCUUAUUGUUCUCCCACUCAAAACUUGGGUCUACUUGAGCCU ((((((.(((((......((((.(....)))))....((...........)).))))))).)))).((((..((...((((.........))))...)).))))... ( -26.40) >DroSec_CAF1 21297 107 + 1 CGGCAGCGAAGGCAACAAUGGAACGAAAGUGCACUCCGCUUUCUUCUGCUGCUCCUGCCUCGCCGUCUUAUUGUUCUCCCACUCGAAACUUGGGUCUACUUGAGCCU .(((.((((.((((.((..((((.(((((((.....)))))))))))..))....)))))))).........((...((((.........))))...))....))). ( -32.40) >DroSim_CAF1 21112 107 + 1 CGGCAGCGAAGGCAACAAUGCAACGAAAGUGCACUCUGCUUUCUUCUGCUGCUCCUGCCUCGCCGUCUUAUUGUUCUCCCACUCGAAACUUGGGUCUACUUGAGCCU ((((((((((((((....((((.(....)))))...)))))))....)))((....))...)))).((((..((...((((.........))))...)).))))... ( -29.40) >DroEre_CAF1 21535 107 + 1 AGGCAGCGAAGGCAGCAAUGCAACGAAAGUGCACUCUGCUUUCUUCUGCUGUUCCUUCCUCUCCGUCUUAUUGUUCUCCCACUCGAAACUUGGGCCUACUUGAGCCU .(((((.((((((((...((((.(....)))))..))))))))..)))))..............(.((((..((...((((.........))))...)).)))).). ( -30.40) >DroYak_CAF1 21560 107 + 1 CGGCAGCGAAGGCAACAAUGCAACGAAAGUGCACUCUGCUUUCUUCGGCUGCUCCUUCCGCGCCGUCUUAUUGUUCUCCCACUCGAAACUUGGGUCUACUUGAGCCU .(((...(((((((....((((.(....)))))...)))))))..((((.((.......)))))).......((...((((.........))))...))....))). ( -31.50) >consensus CGGCAGCGAAGGCAACAAUGCAACGAAAGUGCACUCUGCUUUCUUCUGCUGCUCCUUCCUCGCCGUCUUAUUGUUCUCCCACUCGAAACUUGGGUCUACUUGAGCCU .(((((((((((((....((((.(....)))))...)))))))....))))))...........(.((((..((...((((.........))))...)).)))).). (-25.88 = -26.32 + 0.44)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:39:48 2006