
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 8,284,495 – 8,284,589 |
| Length | 94 |
| Max. P | 0.877287 |

| Location | 8,284,495 – 8,284,589 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 91.47 |
| Mean single sequence MFE | -15.79 |
| Consensus MFE | -11.65 |
| Energy contribution | -11.69 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.769928 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 8284495 94 + 22407834 AUAUAGUUCUGCUCCAAUUCAUUCUAAUUAAAUUCCACCACAAAUGCGGAAGUUCGUUUUCGUUUUCAACAAAC-------AAGAAACAUUUCGAGCA-ACA .........(((((..................((((.(.......).))))....(((((.((((.....))))-------..))))).....)))))-... ( -12.10) >DroSec_CAF1 62298 101 + 1 AUAUAGUUCUGCUCCAAUUCAUUCUAAUUAAAUUCCACCACGAACACGGAAGUUCGUUUUCGUUUUCAACAAACAAGAAACAAGAAACACUUCGAGCA-ACA .........(((((.......((((..............((((((......))))))....((((.....)))).))))....(((....))))))))-... ( -15.50) >DroSim_CAF1 65681 101 + 1 AUAUAGUUCUGCUCCAAUUCAUUCUAAUUAAAUUCCACCACGAACACGGAAGUUCGUUUUCGUUUUCAACAAACGAGAAACAAGAAACACUUCGAGCA-ACA .........(((((.......((((..............((((((......))))))((((((((.....))))))))....)))).......)))))-... ( -19.94) >DroEre_CAF1 64989 95 + 1 AUAUAGUUCUGCUCCAAUUCAUUCUAAUUAAAUUCCACCACGAACACGGAAGUUCGUUUUCGUUUUCAACACAU-------AAGAAACACUCCGAGCAAACA .....(((.(((((.........................((((((......))))))....(((((........-------..))))).....)))))))). ( -15.70) >DroYak_CAF1 66544 95 + 1 AUAUAGUUCUGCUCCAAUUCAUUCUAAUUAAAUUCCACCACGAACACGGAAGUUCGUUUUCGUUUUCAACAACU-------AAGAAACACUCCGAGCAAACA .....(((.(((((.........................((((((......))))))....(((((........-------..))))).....)))))))). ( -15.70) >consensus AUAUAGUUCUGCUCCAAUUCAUUCUAAUUAAAUUCCACCACGAACACGGAAGUUCGUUUUCGUUUUCAACAAAC_______AAGAAACACUUCGAGCA_ACA .........(((((.........................((((((......))))))....(((((.................))))).....))))).... (-11.65 = -11.69 + 0.04)

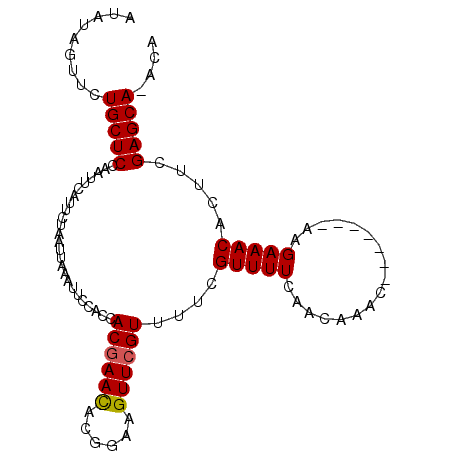

| Location | 8,284,495 – 8,284,589 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 91.47 |
| Mean single sequence MFE | -24.16 |
| Consensus MFE | -19.48 |
| Energy contribution | -19.28 |
| Covariance contribution | -0.20 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.90 |
| SVM RNA-class probability | 0.877287 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 8284495 94 - 22407834 UGU-UGCUCGAAAUGUUUCUU-------GUUUGUUGAAAACGAAAACGAACUUCCGCAUUUGUGGUGGAAUUUAAUUAGAAUGAAUUGGAGCAGAACUAUAU ..(-(((((.((....((((.-------(((((((.........)))))))((((((.......)))))).......))))....)).))))))........ ( -21.90) >DroSec_CAF1 62298 101 - 1 UGU-UGCUCGAAGUGUUUCUUGUUUCUUGUUUGUUGAAAACGAAAACGAACUUCCGUGUUCGUGGUGGAAUUUAAUUAGAAUGAAUUGGAGCAGAACUAUAU ..(-(((((.((....((((.(((((((((((.....))))))..((((((......))))))...)))))......))))....)).))))))........ ( -23.60) >DroSim_CAF1 65681 101 - 1 UGU-UGCUCGAAGUGUUUCUUGUUUCUCGUUUGUUGAAAACGAAAACGAACUUCCGUGUUCGUGGUGGAAUUUAAUUAGAAUGAAUUGGAGCAGAACUAUAU ..(-(((((.((....((((.(((((((((((.....))))))..((((((......))))))...)))))......))))....)).))))))........ ( -25.70) >DroEre_CAF1 64989 95 - 1 UGUUUGCUCGGAGUGUUUCUU-------AUGUGUUGAAAACGAAAACGAACUUCCGUGUUCGUGGUGGAAUUUAAUUAGAAUGAAUUGGAGCAGAACUAUAU ..(((((((.((....((((.-------....((((((..(.(..((((((......))))))..).)..)))))).))))....)).)))))))....... ( -24.80) >DroYak_CAF1 66544 95 - 1 UGUUUGCUCGGAGUGUUUCUU-------AGUUGUUGAAAACGAAAACGAACUUCCGUGUUCGUGGUGGAAUUUAAUUAGAAUGAAUUGGAGCAGAACUAUAU ..(((((((.((....((((.-------....((((((..(.(..((((((......))))))..).)..)))))).))))....)).)))))))....... ( -24.80) >consensus UGU_UGCUCGAAGUGUUUCUU_______GUUUGUUGAAAACGAAAACGAACUUCCGUGUUCGUGGUGGAAUUUAAUUAGAAUGAAUUGGAGCAGAACUAUAU ....(((((.((....((((............((((((..(.(..((((((......))))))..).)..)))))).))))....)).)))))......... (-19.48 = -19.28 + -0.20)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:38:44 2006