
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 8,060,240 – 8,060,347 |
| Length | 107 |
| Max. P | 0.994631 |

| Location | 8,060,240 – 8,060,347 |
|---|---|
| Length | 107 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 74.11 |
| Mean single sequence MFE | -39.58 |
| Consensus MFE | -22.86 |
| Energy contribution | -21.70 |
| Covariance contribution | -1.16 |
| Combinations/Pair | 1.36 |
| Mean z-score | -2.61 |
| Structure conservation index | 0.58 |
| SVM decision value | 2.50 |
| SVM RNA-class probability | 0.994631 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
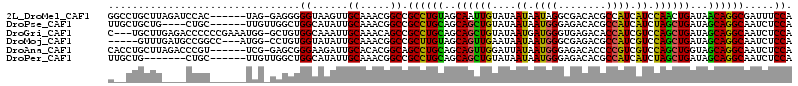
>2L_DroMel_CAF1 8060240 107 + 22407834 GGCCUGCUUAGAUCCAC------UAG-GAGGGGGUAAGUUGCAAACGGCCGCCUGUAGCAAUUGUAUAAUAAUAGGCGACACGCCAUCAUCCAACUGAUAACAGGCGAUUUCCA (((((((....((((.(------(..-..)))))).....)))...))))((((((..................((((...))))((((......)))).))))))........ ( -31.70) >DroPse_CAF1 64469 104 + 1 UUGCUGCUG----CUGC------UUGUUGGCUGGCAUAUUGCAAACGGCCGCCUGCAGCAGCUGUAUAAUAAUGGGAGACACGCCAUCAUCUAGCUGAUAGCAGGCAAUCUCCA .(((.((((----((((------..((.(((((((.....))...)))))))..)))))))).)))........(((((...(((...(((.....)))....)))..))))). ( -40.30) >DroGri_CAF1 19071 110 + 1 C---UGCUUGAGACCCCCCGAAAUGG-GCUGUGGCAAAUUGCAAACAGCCGCCUGCAGCAGCUGUAUAAUGAUGGGUGAGACACCAUCGUCCAGCUGAUAGCAGGCAAUCUCCA .---.....((((............(-(((((.((.....))..))))))((((((..((((((....(((((((........))))))).))))))...))))))..)))).. ( -46.70) >DroMoj_CAF1 17184 105 + 1 -----GUUUGAUGCCGGCC---AUGG-CCUGUGGUAUAUUGCAAACGGCCGCUUGUAGCAGUUGAAUAAUAAUGGGCGAGACGCCAUCGUCCAGCUGAUAGCAGGCAAUCUCCA -----(((((((((((((.---...)-))...)))))....))))).((((((..((((.(((....)))..(((((((.......)))))))))))..))).)))........ ( -33.60) >DroAna_CAF1 18574 107 + 1 CACCUGCUUAGACCCGU------UCG-GAGCGGGAAGAUUGCACACGGCAGCCUGCAGCAGUUGGAUUAUAAUGGGAGACACCCCGUCGUCCAGCUGGUAGCAGGCAAUCUCCA ..(((((((.((.....------)).-))))))).(((((((.....)).(((((((.(((((((((....(((((......))))).))))))))).).)))))))))))... ( -47.10) >DroPer_CAF1 85120 101 + 1 UUGCUG-------CUGC------UUGUUGGCUGGCAUAUUGCAAACGGCCGCCUGCAGCAGCUGUAUAAUAAUGGGAGACACGCCAUCAUCUAGCUGAUAGCAGGCAAUCUCCA ..((((-------((((------..((.(((((((.....))...)))))))..))))))))............(((((...(((...(((.....)))....)))..))))). ( -38.10) >consensus C__CUGCUU____CCGC______UGG_GAGCUGGCAUAUUGCAAACGGCCGCCUGCAGCAGCUGUAUAAUAAUGGGAGACACGCCAUCAUCCAGCUGAUAGCAGGCAAUCUCCA ................................((......((.....)).((((((..((((((....((.((((........)))).)).))))))...)))))).....)). (-22.86 = -21.70 + -1.16)

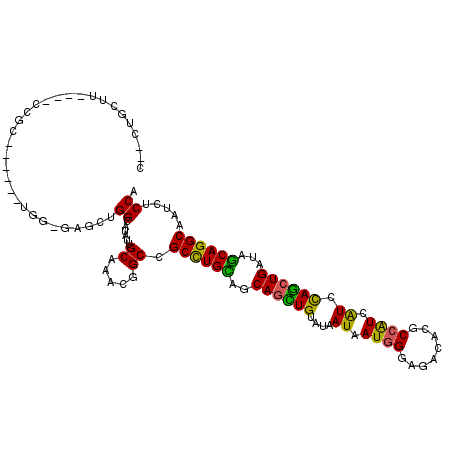

| Location | 8,060,240 – 8,060,347 |
|---|---|
| Length | 107 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 74.11 |
| Mean single sequence MFE | -37.30 |
| Consensus MFE | -21.35 |
| Energy contribution | -21.77 |
| Covariance contribution | 0.42 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.57 |
| SVM decision value | 1.34 |
| SVM RNA-class probability | 0.943680 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 8060240 107 - 22407834 UGGAAAUCGCCUGUUAUCAGUUGGAUGAUGGCGUGUCGCCUAUUAUUAUACAAUUGCUACAGGCGGCCGUUUGCAACUUACCCCCUC-CUA------GUGGAUCUAAGCAGGCC .((...((((((((...((((((((((((((((...)))).))))))...))))))..))))))))))((((((......((.(...-...------).))......)))))). ( -32.80) >DroPse_CAF1 64469 104 - 1 UGGAGAUUGCCUGCUAUCAGCUAGAUGAUGGCGUGUCUCCCAUUAUUAUACAGCUGCUGCAGGCGGCCGUUUGCAAUAUGCCAGCCAACAA------GCAG----CAGCAGCAA .((((((....((((((((......)))))))).))))))............((((((((.((((((.....))....)))).((......------)).)----))))))).. ( -40.80) >DroGri_CAF1 19071 110 - 1 UGGAGAUUGCCUGCUAUCAGCUGGACGAUGGUGUCUCACCCAUCAUUAUACAGCUGCUGCAGGCGGCUGUUUGCAAUUUGCCACAGC-CCAUUUCGGGGGGUCUCAAGCA---G ..((((((((((((...((((((...(((((((.......)))))))...))))))..))))))((((((..((.....)).)))))-)..........)))))).....---. ( -46.50) >DroMoj_CAF1 17184 105 - 1 UGGAGAUUGCCUGCUAUCAGCUGGACGAUGGCGUCUCGCCCAUUAUUAUUCAACUGCUACAAGCGGCCGUUUGCAAUAUACCACAGG-CCAU---GGCCGGCAUCAAAC----- ....(((.(((.((((((((.((((..((((((...))))....))..)))).)))........((((...((........))..))-))))---))).))))))....----- ( -29.40) >DroAna_CAF1 18574 107 - 1 UGGAGAUUGCCUGCUACCAGCUGGACGACGGGGUGUCUCCCAUUAUAAUCCAACUGCUGCAGGCUGCCGUGUGCAAUCUUCCCGCUC-CGA------ACGGGUCUAAGCAGGUG ........((((((...(((.((((.....(((.....))).......)))).)))..)))))).(((.(((........((((...-...------.)))).....)))))). ( -34.12) >DroPer_CAF1 85120 101 - 1 UGGAGAUUGCCUGCUAUCAGCUAGAUGAUGGCGUGUCUCCCAUUAUUAUACAGCUGCUGCAGGCGGCCGUUUGCAAUAUGCCAGCCAACAA------GCAG-------CAGCAA .((((((....((((((((......)))))))).))))))............((((((((.((((((............))).))).....------))))-------)))).. ( -40.20) >consensus UGGAGAUUGCCUGCUAUCAGCUGGACGAUGGCGUGUCUCCCAUUAUUAUACAACUGCUGCAGGCGGCCGUUUGCAAUAUGCCACCCC_CAA______GCGG____AAGCAG__A .((....(((((((...((((((...(((((.(.....))))))......))))))..)))))))((.....))......))................................ (-21.35 = -21.77 + 0.42)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:37:09 2006