
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 7,655,135 – 7,655,240 |
| Length | 105 |
| Max. P | 0.820880 |

| Location | 7,655,135 – 7,655,240 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 79.15 |
| Mean single sequence MFE | -33.83 |
| Consensus MFE | -26.20 |
| Energy contribution | -25.37 |
| Covariance contribution | -0.83 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.820880 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 7655135 105 - 22407834 UGUACC------------CUUCUAUUUGCAGCAAUCGGUCGGCGAGCUCGGAGCAAGAUAAC---UCAGAUCUAUCCGAGCACUCGGAGAAAUCGCCGCUGGUUAGCGCGAGGUUAGACA ......------------..(((((((((.((((((((.((((..(((((((...((((...---....)))).))))))).....((....)))))))))))).)))))))..)))).. ( -39.40) >DroPse_CAF1 1 98 - 1 UCCAUC------------CUUCUAUUUGCAGCAAUCGGUCGGCCAGCUCGGAGCAAGAUAACAUGUCCGACAUGUCCGAGCACUCGGAGAAGUCGCCGCUGGUUAGCGCG---------- ......------------.........((.((((((((.((((......(((.((........)))))(((.(.(((((....))))).).))))))))))))).)))).---------- ( -32.80) >DroGri_CAF1 1 107 - 1 CGAGCGUAUCUCAAUCCUAUUCUAUUUGCAGCAACCGGUCGGCCAGCUCGGAGCAAGAUAAC---UCAGAUCAGUCCGAACACUCGGAGAAAUCGCCACUGGUUAGCGCG---------- .(((.....)))...............((.(((((((((.(((.......(((........)---)).(((...(((((....)))))...))))))))))))).)))).---------- ( -29.50) >DroWil_CAF1 1 89 - 1 ------------------AUUCUAUUUACAGCAAUCGCUCGGCCAGUUCGGAGCAAGAUAAC---UCAGAUCUAUCCGAGCACUCGGAGAAAUCGCCGCUGGUUAGUGCG---------- ------------------............(((...(((((((..(((((((...((((...---....)))).))))))).....((....))))))..)))...))).---------- ( -26.40) >DroMoj_CAF1 1 95 - 1 CGCGUG------------CUUCUAUUUGCAGCAACCGGUCGGCCAGCUCGGAGCAAGAUAAC---UCAGAUCAAUCCGAGCACUCGGAGAAAUCGCCACUGGUUAGCGCG---------- ((((((------------((.(.....).)))(((((((.(((..(((((((....(((...---....)))..))))))).....((....)))))))))))).)))))---------- ( -38.30) >DroPer_CAF1 69 108 - 1 UCCAUC------------CUUCUAUUUGCAGCAAUCGGUCGGCCAGCUCGGAGCAAGAUAACAUGUCCGACAUGUCCGAGCACUCGGAGAAGUCGCCGCUGGUUAGCGCGAGGUUAGACA ......------------..(((((((((.((((((((.((((......(((.((........)))))(((.(.(((((....))))).).))))))))))))).)))))))..)))).. ( -36.60) >consensus UGCAUC____________CUUCUAUUUGCAGCAAUCGGUCGGCCAGCUCGGAGCAAGAUAAC___UCAGAUCUAUCCGAGCACUCGGAGAAAUCGCCGCUGGUUAGCGCG__________ ...........................((.(((((((((.(((..((.....))..............(((...(((((....)))))...))))))))))))).))))........... (-26.20 = -25.37 + -0.83)

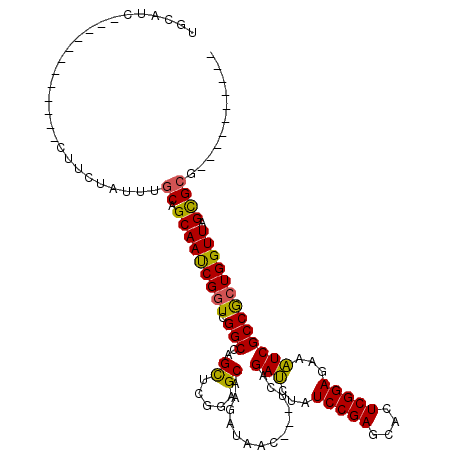
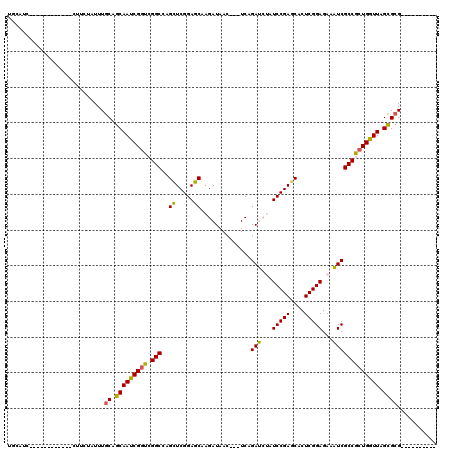
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:34:46 2006