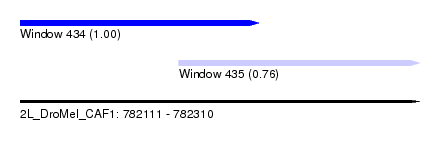
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 782,111 – 782,310 |
| Length | 199 |
| Max. P | 0.997308 |

| Location | 782,111 – 782,230 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.50 |
| Mean single sequence MFE | -26.64 |
| Consensus MFE | -23.34 |
| Energy contribution | -23.94 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.88 |
| SVM decision value | 2.83 |
| SVM RNA-class probability | 0.997308 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
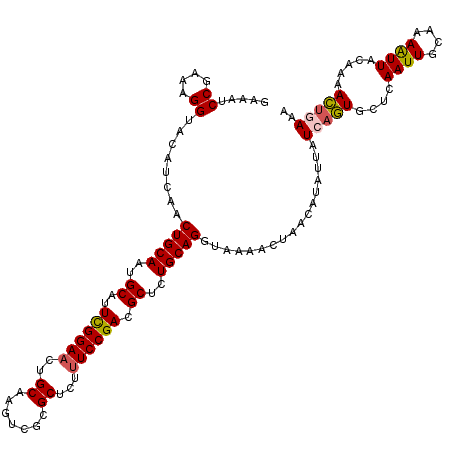
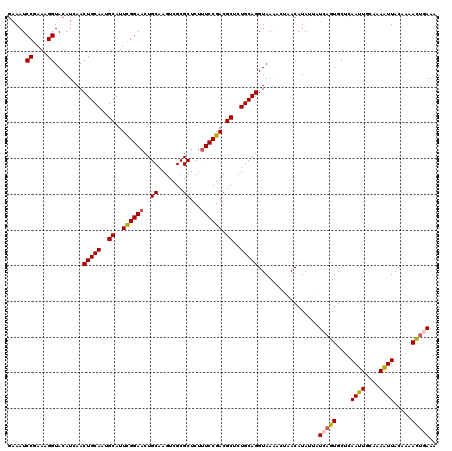
>2L_DroMel_CAF1 782111 119 + 22407834 GAAAUCCGAAAGGUACAUCAACUGCAAUGCAUUCGGAACUGCAAGUCGCGCUCUUUCCGACGCUCUGCAGGUAAAA-UAACAUAUUAUCAGUGCUCAAUUACAAAAUUACUAAACUGAAA ((...((....))....))..(((((..((..((((((..((.......))...)))))).))..)))))......-..........(((((....((((....)))).....))))).. ( -27.30) >DroSec_CAF1 42502 120 + 1 GAAAUCCGAAAGGUACAUCAACUGCAAUGCAUUCGGAACUGCAAGUCGCGCUCUUUCCGACGCUCUGCAGGUAAAAUUAACAUAUAAUCAGUGCUCAAUUGAAAAAUUACAAAACUGAAA ((...((....))....))..(((((..((..((((((..((.......))...)))))).))..))))).................(((((....((((....)))).....))))).. ( -29.70) >DroSim_CAF1 46503 120 + 1 GAAAUCCGAAAGGUACAUCAACUGCAAUGCAUUCGGAACUGCAAGUCGCGCUCUUUCCGACGCUCUGCAGGUAAAAUUAACAUAUAAUCAGUGCUCAAUUGAAAAAUUACUAAACUGAAA ((...((....))....))..(((((..((..((((((..((.......))...)))))).))..))))).................(((((....((((....)))).....))))).. ( -29.70) >DroEre_CAF1 42141 120 + 1 AUAAUCCGAAAGGUACAUCAACUGCAAUGCAUUUGGAACUGCAAGUCGCGCUCUUUCCGACGCUCUGCAGGUAAGACAAACAUAAUUUCAUUGCACAAUUACAAAGUUACAAAAAUUAAG .....((....))........(((((..((..((((((..((.......))...)))))).))..)))))((((((...........)).)))).......................... ( -20.90) >DroYak_CAF1 46108 120 + 1 GAAAUCCGAAAGGUACAUAAACUGCAACGCAUUCGGAACUGCAAGUCGCGCUCUCUCCGACGCUCUGCAGGUAAAACAAACAUAUUUUCAUUGCUAAAUUGCAAAAUUACAAAAACUAAA ((((.((....))........(((((..((..(((((...((.......))....))))).))..)))))...............)))).((((......))))................ ( -25.60) >consensus GAAAUCCGAAAGGUACAUCAACUGCAAUGCAUUCGGAACUGCAAGUCGCGCUCUUUCCGACGCUCUGCAGGUAAAACUAACAUAUUAUCAGUGCUCAAUUGCAAAAUUACAAAACUGAAA .....((....))........(((((..((..((((((..((.......))...)))))).))..))))).................(((((....((((....)))).....))))).. (-23.34 = -23.94 + 0.60)

| Location | 782,190 – 782,310 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.17 |
| Mean single sequence MFE | -29.48 |
| Consensus MFE | -22.32 |
| Energy contribution | -22.56 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.764500 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
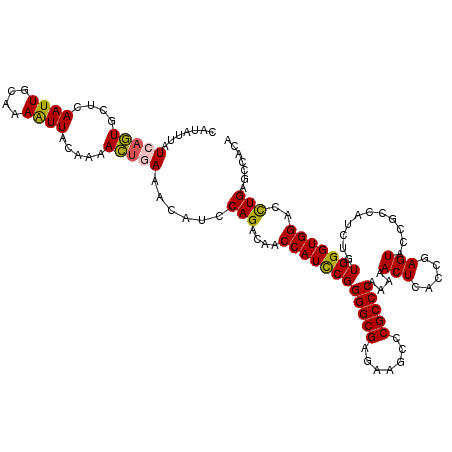
>2L_DroMel_CAF1 782190 120 + 22407834 CAUAUUAUCAGUGCUCAAUUACAAAAUUACUAAACUGAAACAUCCAGACAACCAUCCGGGGCGAGAAGCCCGCCCAAAAACUCACUGAGUACCGCCAUCUAUGGGUGGAUCUGAGCCACA ..........(((((((.................(((.......)))....((((((((((((.......)))))....((((...))))...........)))))))...)))).))). ( -31.40) >DroSec_CAF1 42582 120 + 1 CAUAUAAUCAGUGCUCAAUUGAAAAAUUACAAAACUGAAACAUUCAGACAACCAUCCGGGGCGAAAAGCCCGCCCAAAAACUCACCGAGUACCGCCAUCUGUGGGUGGACCUGAGCCACA ..........(((((((.................((((.....))))....((((((((((((.......)))))....((((...))))...........)))))))...)))).))). ( -32.80) >DroSim_CAF1 46583 120 + 1 CAUAUAAUCAGUGCUCAAUUGAAAAAUUACUAAACUGAAACAAACAGACAACCAUCCGGGGCGAGAAGCCCGCCCAAAAACUCACCGAGUACCGCCAUCUGUGGGUGGACUUGAGUCACA .......(((((....((((....)))).....)))))........(((.........(((((.......)))))..........(((((.(((((.......)))))))))).)))... ( -32.90) >DroEre_CAF1 42221 120 + 1 CAUAAUUUCAUUGCACAAUUACAAAGUUACAAAAAUUAAGCAUCCAGACCACCAUUCGGGGCGAGAAGCCCGCCCAGAAACUGACCGAGUACCGCCAUCUGUGGGUGGACCUGAGUCACA ...........(((..((((.............))))..)))..(((.(((((((((((((((.......))))(((...))).))))))..(((.....))))))))..)))....... ( -25.72) >DroYak_CAF1 46188 120 + 1 CAUAUUUUCAUUGCUAAAUUGCAAAAUUACAAAAACUAAACAUCCAGACAACCAUCCGGGGCGAGAAGCCCGCCCAGAAACUAACCGAGUACCGCCAUCUGUGGGUGGACCUGAGCCACA ..........((((......))))....................(((....((((((((((((.......)))))....(((.....)))...........)))))))..)))....... ( -24.60) >consensus CAUAUUAUCAGUGCUCAAUUGCAAAAUUACAAAACUGAAACAUCCAGACAACCAUCCGGGGCGAGAAGCCCGCCCAAAAACUCACCGAGUACCGCCAUCUGUGGGUGGACCUGAGCCACA .......(((((....((((....)))).....)))))......(((....((((((((((((.......)))))....(((.....)))...........)))))))..)))....... (-22.32 = -22.56 + 0.24)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:31:44 2006