
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 7,630,196 – 7,630,308 |
| Length | 112 |
| Max. P | 0.959506 |

| Location | 7,630,196 – 7,630,308 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 88.10 |
| Mean single sequence MFE | -29.58 |
| Consensus MFE | -22.40 |
| Energy contribution | -23.46 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.79 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.50 |
| SVM RNA-class probability | 0.959506 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
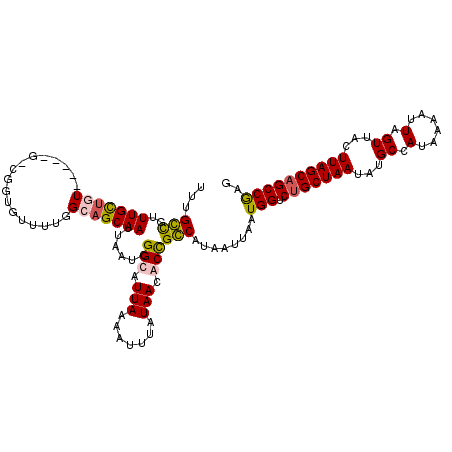
>2L_DroMel_CAF1 7630196 112 + 22407834 CUCGGCUGCUAAGUAACUAAUUUUAUGGCAUAUUAGCAGACCAUUAAUUAUGCCGGUGUUAUAAAUUUUAAUGCCGAUUAAUUGCUGCCAAAACAUAG-C-----AACACAACCAGCG-- ...(((((((((....(((......)))....))))))).)).........((((((((((.......)))))))).....((((((........)))-)-----))........)).-- ( -26.80) >DroPse_CAF1 163693 120 + 1 CUCGGCUGCUAAGUAACUAAUUUUAUGGCAUAUUAGCAGACCAUUAAUUAUGGCAGCGUUAUAAAUUUUAAUGCCGAUUAAUUGCUGCCAAAACACCGGCCAGAAACAGCAAAAGCCAAA ...(((((((((....(((......)))....))))))).))........((((.((((((.......)))))).......((((((((........))).......)))))..)))).. ( -30.81) >DroAna_CAF1 135793 114 + 1 CUUGGCGGCUAAGUAACUAAUUUUAUGGCAUAUUAGCAGACCAUUAAUUAUGCCGGUGUUAUAAAUUUUAAUGCCGAUUAAUUGCUGCCAAAACACGG-C-----AAAGCAAACGGCAAC ..((((.(((((....(((......)))....))))).).))).......(((((((((((.......)))))))......((((((((.......))-)-----..)))))..)))).. ( -29.90) >DroPer_CAF1 159774 120 + 1 CUCGGCUGCUAAGUAACUAAUUUUAUGGCAUAUUAGCAGACCAUUAAUUAUGGCAGCGUUAUAAAUUUUAAUGCCGAUUAAUUGCUGCCAAAACACCGGCCAGAAACAGCAAAAGCCAAA ...(((((((((....(((......)))....))))))).))........((((.((((((.......)))))).......((((((((........))).......)))))..)))).. ( -30.81) >consensus CUCGGCUGCUAAGUAACUAAUUUUAUGGCAUAUUAGCAGACCAUUAAUUAUGCCAGCGUUAUAAAUUUUAAUGCCGAUUAAUUGCUGCCAAAACACCG_C_____AAAGCAAAAGCCAAA ...(((((((((....(((......)))....))))))).))........(((((((((((.......)))))))......((((((...................))))))..)))).. (-22.40 = -23.46 + 1.06)


| Location | 7,630,196 – 7,630,308 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.10 |
| Mean single sequence MFE | -30.39 |
| Consensus MFE | -19.37 |
| Energy contribution | -19.55 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.64 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 7630196 112 - 22407834 --CGCUGGUUGUGUU-----G-CUAUGUUUUGGCAGCAAUUAAUCGGCAUUAAAAUUUAUAACACCGGCAUAAUUAAUGGUCUGCUAAUAUGCCAUAAAAUUAGUUACUUAGCAGCCGAG --.((((((((((((-----(-(((.....))))))))..)))))))).................((((.(((((.(((((.((....)).)))))..)))))(((....))).)))).. ( -30.60) >DroPse_CAF1 163693 120 - 1 UUUGGCUUUUGCUGUUUCUGGCCGGUGUUUUGGCAGCAAUUAAUCGGCAUUAAAAUUUAUAACGCUGCCAUAAUUAAUGGUCUGCUAAUAUGCCAUAAAAUUAGUUACUUAGCAGCCGAG .((((((...((((.....(((((..(((.(((((((..........................))))))).)))...))))).((((((.(......).))))))....)))))))))). ( -31.67) >DroAna_CAF1 135793 114 - 1 GUUGCCGUUUGCUUU-----G-CCGUGUUUUGGCAGCAAUUAAUCGGCAUUAAAAUUUAUAACACCGGCAUAAUUAAUGGUCUGCUAAUAUGCCAUAAAAUUAGUUACUUAGCCGCCAAG ..(((((.(((((..-----(-(((.....))))))))).....)))))................((((.((((((((.....((......))......))))))))....))))..... ( -27.60) >DroPer_CAF1 159774 120 - 1 UUUGGCUUUUGCUGUUUCUGGCCGGUGUUUUGGCAGCAAUUAAUCGGCAUUAAAAUUUAUAACGCUGCCAUAAUUAAUGGUCUGCUAAUAUGCCAUAAAAUUAGUUACUUAGCAGCCGAG .((((((...((((.....(((((..(((.(((((((..........................))))))).)))...))))).((((((.(......).))))))....)))))))))). ( -31.67) >consensus UUUGCCGUUUGCUGU_____G_CGGUGUUUUGGCAGCAAUUAAUCGGCAUUAAAAUUUAUAACACCGCCAUAAUUAAUGGUCUGCUAAUAUGCCAUAAAAUUAGUUACUUAGCAGCCGAG ...(((..(((((((.................)))))))......(((.(((.......))).))))))........(((.(((((((...((.(......).))...)))))))))).. (-19.37 = -19.55 + 0.19)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:34:35 2006