
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 7,495,504 – 7,495,612 |
| Length | 108 |
| Max. P | 0.940316 |

| Location | 7,495,504 – 7,495,612 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.51 |
| Mean single sequence MFE | -31.29 |
| Consensus MFE | -17.12 |
| Energy contribution | -18.65 |
| Covariance contribution | 1.53 |
| Combinations/Pair | 1.08 |
| Mean z-score | -3.04 |
| Structure conservation index | 0.55 |
| SVM decision value | 1.29 |
| SVM RNA-class probability | 0.940243 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 7495504 108 + 22407834 -GAACUUACCAAAACCGCGCUCUCUCGCACCAGCUGAGAGACAUUCUCUCUCAUUUGCACCGCUGAGCCGAGCUUAUUUAAGCCGCGCGUAAACUUUCGCUCCAUCCUG----------- -...............((((((...((...(((.(((((((.....))))))).)))...))..)))).(.((((....)))))))(((........))).........----------- ( -25.70) >DroPse_CAF1 25309 108 + 1 AUUACUUACCAAAACGGCACUCUCACGCACCAGCUGAGAGACAUUCUCUCUCAUCUGGCCCGCCGAAGUUAGCUUAUUCAAGCCGCGCGUAAAUUUACGCUCCAUUCU------------ ..............((((.......((..((((.(((((((.....))))))).))))..))...(((....)))......)))).(((((....)))))........------------ ( -29.80) >DroGri_CAF1 21157 119 + 1 -AAACUUACCAAAACUGCGCUCUCACGCACAACCUGAGAGACGCUCUCUCUCAGCUGUGCGGCCAUGCCGGGCUUAUUCAAGCCCCGCGUGAAUUUACGCUCCAUUCUCAAUUUCCCUUC -...............(((......((((((..((((((((.....)))))))).))))))..(((((.((((((....)))))).)))))......))).................... ( -41.50) >DroEre_CAF1 18591 108 + 1 -GUACUUACCAAAACCGCGCUCUCCCGCACCAGCUGAGAGACAUUCUCUCUCAUUUGCACCGCAGAGCCGAGCUUAUUUAAGCCGCGCGUAAAUUUCCGUUCCAUCCUG----------- -............((((((((((..((...(((.(((((((.....))))))).)))...)).))))).(.((((....)))))))).))...................----------- ( -25.50) >DroYak_CAF1 18728 108 + 1 -GAGCUUACCAAAAUCGCGCUCUCGCGCACCAGCUGAGAGACAUUCUCUCGCAUUUGCACCGCUGAGCCGAGCUUGUUUAAGCCGCGCGUAAACUUUCGCUCCAUCCUG----------- -((((..........(((((..(((.((..((((.(((((.....)))))((....))...)))).)))))((((....)))).))))).........)))).......----------- ( -35.41) >DroPer_CAF1 23583 105 + 1 AUUACUUACCAAAACGGCACUCUCACGCACCAGCUGAGAGACAUUCUCUCUCAUCUGGCCCGCCGAAGUUAGCUUAUUCAAGCCGCGCGUAAAUUUACGCUCCAU--------------- ..............((((.......((..((((.(((((((.....))))))).))))..))...(((....)))......)))).(((((....))))).....--------------- ( -29.80) >consensus _GAACUUACCAAAACCGCGCUCUCACGCACCAGCUGAGAGACAUUCUCUCUCAUCUGCACCGCCGAGCCGAGCUUAUUCAAGCCGCGCGUAAAUUUACGCUCCAUCCUG___________ ................((((((...((...(((.(((((((.....))))))).)))...))..))))...((((....)))).))(((........))).................... (-17.12 = -18.65 + 1.53)



| Location | 7,495,504 – 7,495,612 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.51 |
| Mean single sequence MFE | -41.32 |
| Consensus MFE | -22.67 |
| Energy contribution | -22.28 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.28 |
| Mean z-score | -3.03 |
| Structure conservation index | 0.55 |
| SVM decision value | 1.29 |
| SVM RNA-class probability | 0.940316 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 7495504 108 - 22407834 -----------CAGGAUGGAGCGAAAGUUUACGCGCGGCUUAAAUAAGCUCGGCUCAGCGGUGCAAAUGAGAGAGAAUGUCUCUCAGCUGGUGCGAGAGAGCGCGGUUUUGGUAAGUUC- -----------((((((.((((....)))).((((((((((....)))))..(((((((........(((((((.....)))))))))))).))......)))))))))))........- ( -37.20) >DroPse_CAF1 25309 108 - 1 ------------AGAAUGGAGCGUAAAUUUACGCGCGGCUUGAAUAAGCUAACUUCGGCGGGCCAGAUGAGAGAGAAUGUCUCUCAGCUGGUGCGUGAGAGUGCCGUUUUGGUAAGUAAU ------------(((((((.(((((....)))))..(((((....))))).((((..(((.(((((.(((((((.....))))))).))))).)))..)))).))))))).......... ( -42.20) >DroGri_CAF1 21157 119 - 1 GAAGGGAAAUUGAGAAUGGAGCGUAAAUUCACGCGGGGCUUGAAUAAGCCCGGCAUGGCCGCACAGCUGAGAGAGAGCGUCUCUCAGGUUGUGCGUGAGAGCGCAGUUUUGGUAAGUUU- ........((..(((.....((((...(((((((.((((((....)))))).))......((((((((((((((.....))))))).)))))))))))).))))..)))..))......- ( -48.00) >DroEre_CAF1 18591 108 - 1 -----------CAGGAUGGAACGGAAAUUUACGCGCGGCUUAAAUAAGCUCGGCUCUGCGGUGCAAAUGAGAGAGAAUGUCUCUCAGCUGGUGCGGGAGAGCGCGGUUUUGGUAAGUAC- -----------.........((.(((((...(((.((((((....)))).))(((((.(.(..(...(((((((.....)))))))....)..).).))))))))))))).))......- ( -38.30) >DroYak_CAF1 18728 108 - 1 -----------CAGGAUGGAGCGAAAGUUUACGCGCGGCUUAAACAAGCUCGGCUCAGCGGUGCAAAUGCGAGAGAAUGUCUCUCAGCUGGUGCGCGAGAGCGCGAUUUUGGUAAGCUC- -----------((((((.((((....)))).(((((.((((....))))((((((((((...((....))(((((.....))))).))))).)).)))..)))))))))))........- ( -41.90) >DroPer_CAF1 23583 105 - 1 ---------------AUGGAGCGUAAAUUUACGCGCGGCUUGAAUAAGCUAACUUCGGCGGGCCAGAUGAGAGAGAAUGUCUCUCAGCUGGUGCGUGAGAGUGCCGUUUUGGUAAGUAAU ---------------.....(((((....)))))...(((((...((((..((((..(((.(((((.(((((((.....))))))).))))).)))..))))...))))...)))))... ( -40.30) >consensus ___________CAGGAUGGAGCGUAAAUUUACGCGCGGCUUAAAUAAGCUCGGCUCGGCGGUGCAAAUGAGAGAGAAUGUCUCUCAGCUGGUGCGUGAGAGCGCGGUUUUGGUAAGUAC_ ..........................((((((((.(((((.......(((......)))...........(((((.....))))))))))))((((....)))).......))))))... (-22.67 = -22.28 + -0.38)

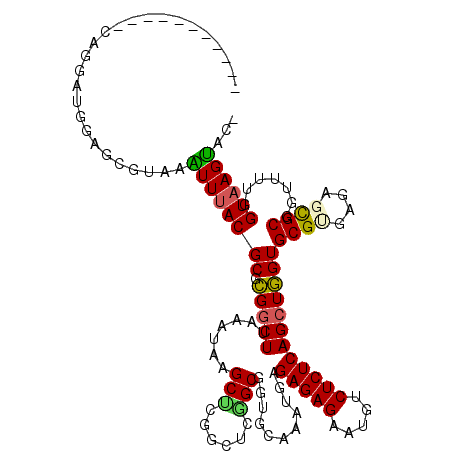

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:33:28 2006