
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 7,313,168 – 7,313,278 |
| Length | 110 |
| Max. P | 0.640703 |

| Location | 7,313,168 – 7,313,278 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 75.46 |
| Mean single sequence MFE | -22.21 |
| Consensus MFE | -11.50 |
| Energy contribution | -12.06 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.640703 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 7313168 110 + 22407834 U-GAGAAACGAAUA---CAAAUAGAGCCAAAAUACUACUAUACAAUACUAUAAGAAAUGAGGUAUGAUUUGGCUUUAUUUCAUAUGCUCUUUU--CGCAGUGUACCUCGUAAUAAG .-.....(((((((---(((((((((((((((((((.(....................).)))))..)))))))))))))....(((......--.)))))))...))))...... ( -22.05) >DroSec_CAF1 106066 103 + 1 U-GGGAAACGGAUAUUCUAAAUAGAGCCAAUA----------UAAUACUAUACGAAAUGAGGUAUUAUUUGGCUUUAUUUCGUAUACUCUUUU--CGCAGUGUACCUCGUAAUAAG (-(.((((.(..(((.(.((((((((((((.(----------(((((((...........)))))))))))))))))))).).)))..).)))--).))................. ( -27.20) >DroSim_CAF1 112931 100 + 1 U-GAGAAACGGAUA---UAAAUAGAGCCAAUU----------UAAUCCUAUACGAAAUAAGGUAUGAUUUGGCUUUAUUUCAUAUACUCUUUU--CGCAGUGUACCUCGUAAUAAG .-(((.........---.((((((((((((.(----------((..(((..........)))..))).))))))))))))..((((((.....--...)))))).)))........ ( -21.90) >DroYak_CAF1 109979 92 + 1 UGAAAAAACGUUUA---UAAAUAGAGUCU--------------------AUAC-AAAUAUGGUAUUUUAUGGCUCUAUUUUAUAUACCUUUUUUGCUCAGUGCACCUCGUAAUAAG ..((((((.((.((---(((((((((((.--------------------((((-.......)))).....))))))).)))))).)).))))))((.....))............. ( -17.70) >consensus U_GAGAAACGGAUA___UAAAUAGAGCCAAUA__________UAAUACUAUACGAAAUAAGGUAUGAUUUGGCUUUAUUUCAUAUACUCUUUU__CGCAGUGUACCUCGUAAUAAG .......((((.......((((((((((((......................................))))))))))))..((((((..........))))))..))))...... (-11.50 = -12.06 + 0.56)

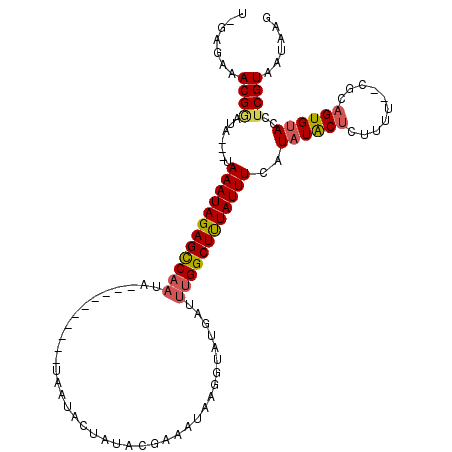

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:32:12 2006