| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 7,211,145 – 7,211,238 |
| Length | 93 |
| Max. P | 0.999479 |

| Location | 7,211,145 – 7,211,238 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 84.70 |
| Mean single sequence MFE | -29.70 |
| Consensus MFE | -24.16 |
| Energy contribution | -24.00 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.14 |
| Mean z-score | -4.17 |
| Structure conservation index | 0.81 |
| SVM decision value | 3.64 |
| SVM RNA-class probability | 0.999479 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 7211145 93 + 22407834 UACAAACAGGCAUGAAUACAUGUACAUAUACAAUAUAUGUAUGUAGCAUGCCGCAUUUGGCGGCGUGCCUAAGCCUGCAAUGUAAACAAUUUU ((((..(((((........(((((((((((...))))))))))).(((((((((.....)))))))))....)))))...))))......... ( -35.90) >DroSec_CAF1 5201 90 + 1 UACAAACAGGCAUGAAUACAUAUGGAUGUAC---AUAUACAUGUAGCAUGCCGCAUUUGGCGGCGUGCCUGAGCCUGCAAUGUAAACAAUUUU ((((..(((((...........((.(((((.---...))))).))(((((((((.....)))))))))....)))))...))))......... ( -31.90) >DroSim_CAF1 5239 90 + 1 UACAAACAGGCAUGAAUACAUAUGGAUGUAC---AUAUAUAUGUAGCAUGCCGCAUUUGGCGGCGUGCCUGAGCCUGCAAUGUAAACAAUUUU ((((..(((((.....((((((((.((....---)).))))))))(((((((((.....)))))))))....)))))...))))......... ( -34.40) >DroEre_CAF1 5233 80 + 1 UACAAACAGGCAUGAACACAUAU-------------ACGAGUAUACCGUGCCGCAUUUGGCGGUGUGCCUGAGCCUGCAAUGUAAACAAUUUU ((((..((((((((....)))..-------------.((.(((((((..((((....))))))))))).)).)))))...))))......... ( -23.50) >DroYak_CAF1 5189 80 + 1 UACAAACAGGCAUGAAUACAUAU-------------ACGAAUGUACUAUGCCGCAUUUGGCGGUGUGCCUGAGCCUGCAAUGUAAACAAUUUU ((((..(((((.....(((((..-------------....))))).((((((((.....)))))))).....)))))...))))......... ( -22.80) >consensus UACAAACAGGCAUGAAUACAUAU__AU_UAC___AUAUGAAUGUAGCAUGCCGCAUUUGGCGGCGUGCCUGAGCCUGCAAUGUAAACAAUUUU ((((..((((((((....)))........................(((((((((.....)))))))))....)))))...))))......... (-24.16 = -24.00 + -0.16)


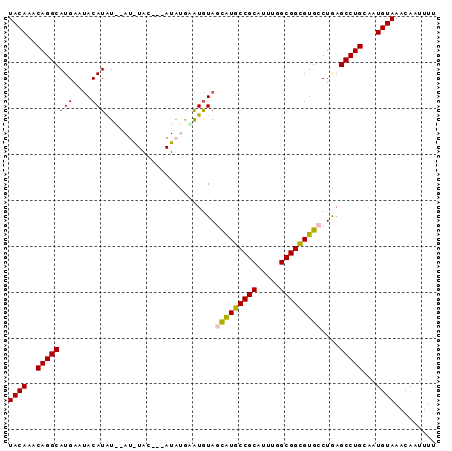
| Location | 7,211,145 – 7,211,238 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 84.70 |
| Mean single sequence MFE | -29.00 |
| Consensus MFE | -20.98 |
| Energy contribution | -22.18 |
| Covariance contribution | 1.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -4.10 |
| Structure conservation index | 0.72 |
| SVM decision value | 2.99 |
| SVM RNA-class probability | 0.998027 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 7211145 93 - 22407834 AAAAUUGUUUACAUUGCAGGCUUAGGCACGCCGCCAAAUGCGGCAUGCUACAUACAUAUAUUGUAUAUGUACAUGUAUUCAUGCCUGUUUGUA .........((((..((((((...((((.(((((.....))))).))))(((((((((((...))))))))..)))......)))))).)))) ( -32.40) >DroSec_CAF1 5201 90 - 1 AAAAUUGUUUACAUUGCAGGCUCAGGCACGCCGCCAAAUGCGGCAUGCUACAUGUAUAU---GUACAUCCAUAUGUAUUCAUGCCUGUUUGUA .........((((..((((((...((((.(((((.....))))).))))..(((.((((---(((......))))))).))))))))).)))) ( -30.80) >DroSim_CAF1 5239 90 - 1 AAAAUUGUUUACAUUGCAGGCUCAGGCACGCCGCCAAAUGCGGCAUGCUACAUAUAUAU---GUACAUCCAUAUGUAUUCAUGCCUGUUUGUA .........((((..((((((...((((.(((((.....))))).))))..((((((((---(......)))))))))....)))))).)))) ( -33.20) >DroEre_CAF1 5233 80 - 1 AAAAUUGUUUACAUUGCAGGCUCAGGCACACCGCCAAAUGCGGCACGGUAUACUCGU-------------AUAUGUGUUCAUGCCUGUUUGUA .........((((..((((((...((((((((((.....)))).((((.....))))-------------...))))))...)))))).)))) ( -25.60) >DroYak_CAF1 5189 80 - 1 AAAAUUGUUUACAUUGCAGGCUCAGGCACACCGCCAAAUGCGGCAUAGUACAUUCGU-------------AUAUGUAUUCAUGCCUGUUUGUA ..............(((((((...(((.....)))......((((((((((((....-------------..))))))).))))).))))))) ( -23.00) >consensus AAAAUUGUUUACAUUGCAGGCUCAGGCACGCCGCCAAAUGCGGCAUGCUACAUACAUAU___GUA_AU__AUAUGUAUUCAUGCCUGUUUGUA .........((((..((((((...((((.(((((.....))))).)))).......................(((....))))))))).)))) (-20.98 = -22.18 + 1.20)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:31:49 2006