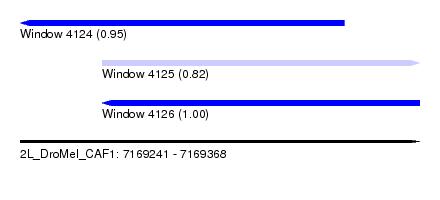

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 7,169,241 – 7,169,368 |
| Length | 127 |
| Max. P | 0.996756 |

| Location | 7,169,241 – 7,169,344 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.58 |
| Mean single sequence MFE | -29.82 |
| Consensus MFE | -18.91 |
| Energy contribution | -19.47 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.04 |
| Mean z-score | -3.01 |
| Structure conservation index | 0.63 |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.952304 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 7169241 103 - 22407834 GGCAGUUUUUAAUUGCGCCAUAAAUCAAAUCUACG--AUAAAUGG-CAAGCAGUUAAAGGGCUGCUAACCAAAUGGGAGCCAUAAGCUGAUAACAGACU--------------GUGUAUA .((((((((((((((((((((..(((........)--))..))))-)..))))))))).((((.(((......))).))))..............))))--------------))..... ( -31.30) >DroSec_CAF1 110517 103 - 1 GGCAGUUUUUAAUUGCGCCAUAAAUCAAAUCUACG--AUAAAUGG-CAAGCAGUUAAAGGGCUGCUAACCAAAUGGGAGCCAUAAGCUGAUAACAGACU--------------GUGUAUA .((((((((((((((((((((..(((........)--))..))))-)..))))))))).((((.(((......))).))))..............))))--------------))..... ( -31.30) >DroWil_CAF1 185272 116 - 1 ---GAUUUUUAAUUGCGCCAUAAAUCAAAUCUAGGAGAUAAAUGGGCCAACAGUUAAAGGGCUGCUAACCAAAUGGGAGCCAUAAGCUGAUAAGAGAGAGAGAGCAACUGACUUUUCA-G ---..((((((...((.((((..(((..........)))..))))))...(((((....((((.(((......))).))))...))))).)))))).(((((..(....)..))))).-. ( -24.00) >DroAna_CAF1 116593 101 - 1 GGCAGUUUUUAAUUGCGCCAUAAAUCAAAUCUACG--AUAAAUGG-CA-GCAGUUAAAGGGCUGCUAACCAAAUGGGAGCCAUAAGCUGAUAACAGACU--------------GUGUA-A .((((((((((((((((((((..(((........)--))..))))-).-))))))))).((((.(((......))).))))..............))))--------------))...-. ( -32.70) >consensus GGCAGUUUUUAAUUGCGCCAUAAAUCAAAUCUACG__AUAAAUGG_CAAGCAGUUAAAGGGCUGCUAACCAAAUGGGAGCCAUAAGCUGAUAACAGACU______________GUGUA_A ..((((((((((((((.((((....................))))....))))))))).((((.(((......))).))))...)))))............................... (-18.91 = -19.47 + 0.56)


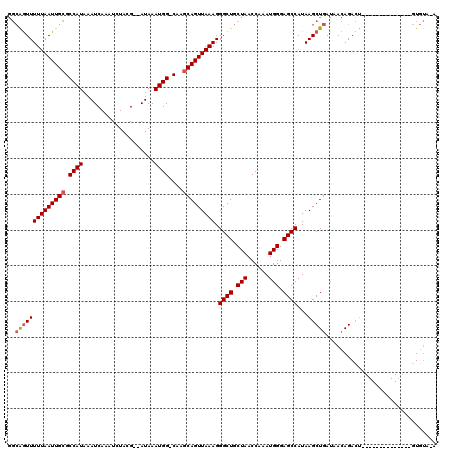
| Location | 7,169,267 – 7,169,368 |
|---|---|
| Length | 101 |
| Sequences | 3 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 86.87 |
| Mean single sequence MFE | -34.37 |
| Consensus MFE | -28.42 |
| Energy contribution | -28.53 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.83 |
| SVM decision value | 0.67 |
| SVM RNA-class probability | 0.818556 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 7169267 101 + 22407834 GCUCCCAUUUGGUUAGCAGCCCUUUAACUGCUUGCCAUUUAUCGUAGAUUUGAUUUAUGGCGCAAUUAAAAACUGCCCCAGAUGAAAA-------------GUGGGGGGGG---GGU ((((((.((((((((((((........))))).(((((..((((......))))..)))))..))))))).....(((((..(....)-------------.))))).)))---))) ( -36.00) >DroSec_CAF1 110543 101 + 1 GCUCCCAUUUGGUUAGCAGCCCUUUAACUGCUUGCCAUUUAUCGUAGAUUUGAUUUAUGGCGCAAUUAAAAACUGCCCCAGAUGAAAA-------------GUGGGCGGAG---GUU .(((((((((((...((((...(((((.(((..(((((..((((......))))..)))))))).)))))..)))).)))))))....-------------(....)))))---... ( -32.50) >DroAna_CAF1 116618 116 + 1 GCUCCCAUUUGGUUAGCAGCCCUUUAACUGC-UGCCAUUUAUCGUAGAUUUGAUUUAUGGCGCAAUUAAAAACUGCCCCAGAUGAAAAAGACGGCGAUGGUGUGGAGGGACGAUGGU .(((((((((((...((((...(((((.(((-.(((((..((((......))))..)))))))).)))))..)))).)))))))......(((.(...).))))))).......... ( -34.60) >consensus GCUCCCAUUUGGUUAGCAGCCCUUUAACUGCUUGCCAUUUAUCGUAGAUUUGAUUUAUGGCGCAAUUAAAAACUGCCCCAGAUGAAAA_____________GUGGGGGGAG___GGU .(((((((((((...((((...(((((.(((..(((((..((((......))))..)))))))).)))))..)))).)))))))...................)))).......... (-28.42 = -28.53 + 0.11)


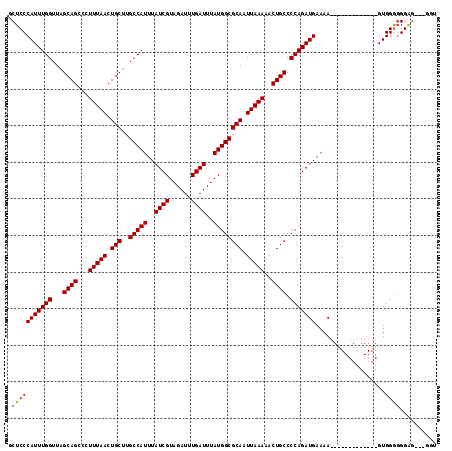
| Location | 7,169,267 – 7,169,368 |
|---|---|
| Length | 101 |
| Sequences | 3 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 86.87 |
| Mean single sequence MFE | -33.30 |
| Consensus MFE | -32.47 |
| Energy contribution | -32.47 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.36 |
| Structure conservation index | 0.97 |
| SVM decision value | 2.74 |
| SVM RNA-class probability | 0.996756 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 7169267 101 - 22407834 ACC---CCCCCCCCAC-------------UUUUCAUCUGGGGCAGUUUUUAAUUGCGCCAUAAAUCAAAUCUACGAUAAAUGGCAAGCAGUUAAAGGGCUGCUAACCAAAUGGGAGC ...---..........-------------((..(((.((((((((((((((((((((((((..(((........)))..)))))..))))))))).)))))))..))).)))..)). ( -32.50) >DroSec_CAF1 110543 101 - 1 AAC---CUCCGCCCAC-------------UUUUCAUCUGGGGCAGUUUUUAAUUGCGCCAUAAAUCAAAUCUACGAUAAAUGGCAAGCAGUUAAAGGGCUGCUAACCAAAUGGGAGC ...---((((.((((.-------------........))))((((((((((((((((((((..(((........)))..)))))..))))))))).))))))..........)))). ( -33.40) >DroAna_CAF1 116618 116 - 1 ACCAUCGUCCCUCCACACCAUCGCCGUCUUUUUCAUCUGGGGCAGUUUUUAAUUGCGCCAUAAAUCAAAUCUACGAUAAAUGGCA-GCAGUUAAAGGGCUGCUAACCAAAUGGGAGC .............................((..(((.((((((((((((((((((((((((..(((........)))..))))).-))))))))).)))))))..))).)))..)). ( -34.00) >consensus ACC___CUCCCCCCAC_____________UUUUCAUCUGGGGCAGUUUUUAAUUGCGCCAUAAAUCAAAUCUACGAUAAAUGGCAAGCAGUUAAAGGGCUGCUAACCAAAUGGGAGC .............................((..(((.((((((((((((((((((((((((..(((........)))..)))))..))))))))).)))))))..))).)))..)). (-32.47 = -32.47 + -0.00)
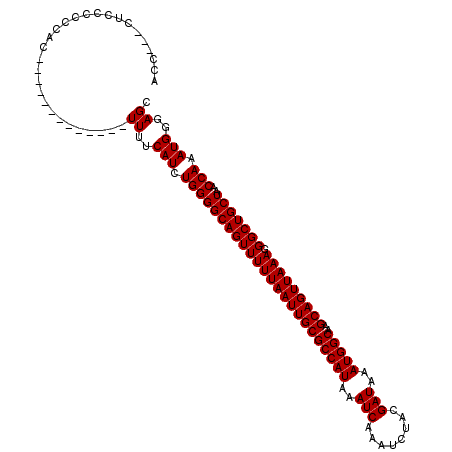


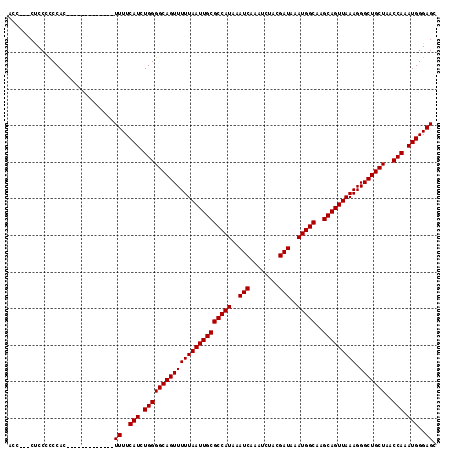
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:31:20 2006