
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 7,070,015 – 7,070,109 |
| Length | 94 |
| Max. P | 0.999670 |

| Location | 7,070,015 – 7,070,109 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 74.19 |
| Mean single sequence MFE | -21.76 |
| Consensus MFE | -5.79 |
| Energy contribution | -8.99 |
| Covariance contribution | 3.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.27 |
| SVM decision value | 1.98 |
| SVM RNA-class probability | 0.984534 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 7070015 94 + 22407834 UUAUCCGAACGAACAGUUCGGAUUUCUGGAUUUCUUUGUUCUAUGGCCAUUGCGUAUAUUUAAACACAGUUUUUCUUCAAAUGCCAUGAACUUU-------------------------- ..((((((((.....))))))))..............((((.(((((.((((.((........)).))))............)))))))))...-------------------------- ( -21.12) >DroSec_CAF1 11886 93 + 1 UUAUCCGAACGAUCAGUUCGGAUUUCUGGAUUUCUUUGUUCUAUGGCCAUUGCGUAUCUUUAAACGCAGUUUUUCUUCA-AUGCCAUGAACUUU-------------------------- ..((((((((.....))))))))..............((((.(((((.(((((((........))))))).........-..)))))))))...-------------------------- ( -26.60) >DroSim_CAF1 12042 93 + 1 UAAUCCGAACGAUCAGUUCGGAUUUCUGGAUUUCUUUGUUCUAUGGCCAUUGCGUAUCUUUAAACGCAGUUUUUCCUUA-AUGCCAUGAACUUU-------------------------- .(((((((((.....))))))))).............((((.(((((.(((((((........))))))).........-..)))))))))...-------------------------- ( -27.50) >DroEre_CAF1 15650 116 + 1 UCAUCCAAACGAUCAGUUA---GUUCGGGAUUCCUUUGUUCUAUGGCCAUCGCGUAUCCCUAACCACAGUUUUACUUCC-AUAGCACGAACUUCCAUCAUUACUUGGGAUAGAAACUUUU ((((((...(((.(.....---).)))(((....(((((.(((((((....))(((...((......))...)))...)-)))).)))))..)))...........)))).))....... ( -18.80) >DroYak_CAF1 15490 91 + 1 AUAUCCAAACGAUCAG-------UUCUGGAUUUCUUUGUUCUAUGGCCAUCGCGUAUCUUUAACCACAGUGUUACUAGC-AUAGCACGAACUUUCAUCC--------------------- ..((((((((.....)-------)).)))))......((((....((....))...............((((((.....-.))))))))))........--------------------- ( -14.80) >consensus UUAUCCGAACGAUCAGUUCGGAUUUCUGGAUUUCUUUGUUCUAUGGCCAUUGCGUAUCUUUAAACACAGUUUUUCUUCA_AUGCCAUGAACUUU__________________________ ...(((((((.....)))))))...............((((.(((((...................................)))))))))............................. ( -5.79 = -8.99 + 3.20)


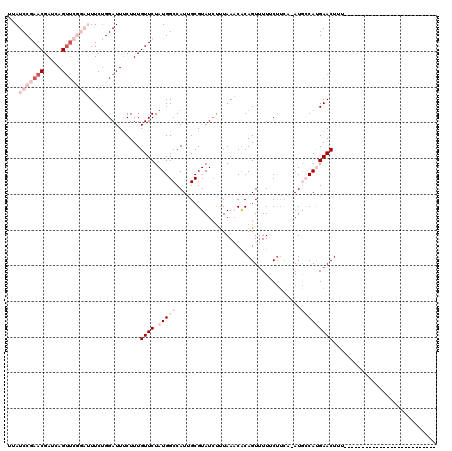
| Location | 7,070,015 – 7,070,109 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 74.19 |
| Mean single sequence MFE | -25.76 |
| Consensus MFE | -13.28 |
| Energy contribution | -15.60 |
| Covariance contribution | 2.32 |
| Combinations/Pair | 1.17 |
| Mean z-score | -3.06 |
| Structure conservation index | 0.52 |
| SVM decision value | 3.86 |
| SVM RNA-class probability | 0.999670 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 7070015 94 - 22407834 --------------------------AAAGUUCAUGGCAUUUGAAGAAAAACUGUGUUUAAAUAUACGCAAUGGCCAUAGAACAAAGAAAUCCAGAAAUCCGAACUGUUCGUUCGGAUAA --------------------------...(((((((((.(((.....)))..(((((........)))))...))))).))))..............((((((((.....)))))))).. ( -25.70) >DroSec_CAF1 11886 93 - 1 --------------------------AAAGUUCAUGGCAU-UGAAGAAAAACUGCGUUUAAAGAUACGCAAUGGCCAUAGAACAAAGAAAUCCAGAAAUCCGAACUGAUCGUUCGGAUAA --------------------------...(((((((((.(-(......))..(((((........)))))...))))).))))..............((((((((.....)))))))).. ( -26.90) >DroSim_CAF1 12042 93 - 1 --------------------------AAAGUUCAUGGCAU-UAAGGAAAAACUGCGUUUAAAGAUACGCAAUGGCCAUAGAACAAAGAAAUCCAGAAAUCCGAACUGAUCGUUCGGAUUA --------------------------...(((((((((..-..((......))((((........))))....))))).)))).............(((((((((.....))))))))). ( -29.10) >DroEre_CAF1 15650 116 - 1 AAAAGUUUCUAUCCCAAGUAAUGAUGGAAGUUCGUGCUAU-GGAAGUAAAACUGUGGUUAGGGAUACGCGAUGGCCAUAGAACAAAGGAAUCCCGAAC---UAACUGAUCGUUUGGAUGA .............(((((..((.(.(..((((((((((..-...))))...(((((((((..(.....)..))))))))).............)))))---)..)).))..))))).... ( -26.10) >DroYak_CAF1 15490 91 - 1 ---------------------GGAUGAAAGUUCGUGCUAU-GCUAGUAACACUGUGGUUAAAGAUACGCGAUGGCCAUAGAACAAAGAAAUCCAGAA-------CUGAUCGUUUGGAUAU ---------------------((((((.(((((.(((((.-..)))))...(((((((((..(.....)..)))))))))..............)))-------))..))))))...... ( -21.00) >consensus __________________________AAAGUUCAUGGCAU_UGAAGAAAAACUGUGUUUAAAGAUACGCAAUGGCCAUAGAACAAAGAAAUCCAGAAAUCCGAACUGAUCGUUCGGAUAA .............................(((((((((......((.....))((((........))))....))))).))))..............((((((((.....)))))))).. (-13.28 = -15.60 + 2.32)

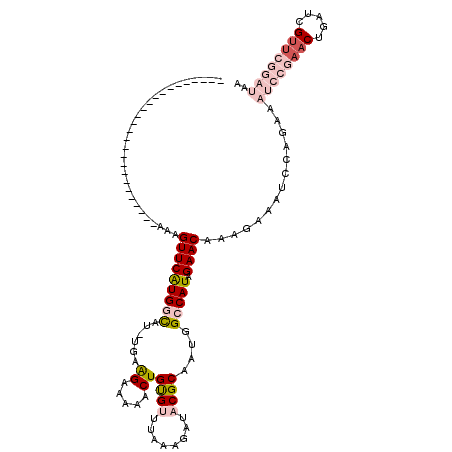

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:30:44 2006