
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 7,006,009 – 7,006,121 |
| Length | 112 |
| Max. P | 0.500000 |

| Location | 7,006,009 – 7,006,121 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.76 |
| Mean single sequence MFE | -22.40 |
| Consensus MFE | -13.72 |
| Energy contribution | -12.28 |
| Covariance contribution | -1.44 |
| Combinations/Pair | 1.38 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.61 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
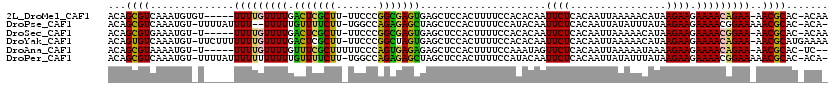
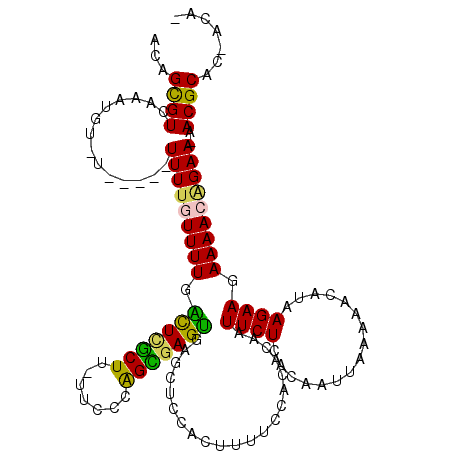
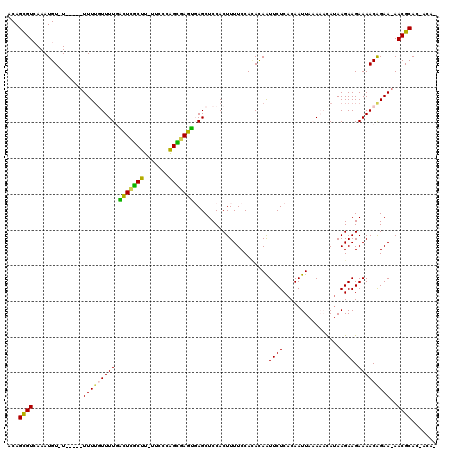
>2L_DroMel_CAF1 7006009 112 + 22407834 ACAGCGUCAAAUGUGU-----UUUUGUUUUGACUCGCUU-UUCCCGGCGAGUGAGCUCCACUUUUCCACACAAUUCUCACAAUUAAAAACAUAAGAAGAAAACAGAA-AACGCAC-ACAA ...((((....(((.(-----((((.((.((..(((((.-.....)))))(((((....................))))).........)).)).))))).)))...-.))))..-.... ( -21.85) >DroPse_CAF1 36224 114 + 1 ACAGCGUCAAAUGU-UUUUAUUUU--UUUUUGUUUUCUU-UGGCCAGAGAGCUAGCUCCACUUUUCCAUACAAUUCUCACAAUUAUAUUUAUAAGAAGAAAACGGAAAAACGCAC-ACA- ...((((.(((((.-...)))))(--(((((((((((((-(((..((.(((....)))..))...))...............((((....)))))))))))))))))))))))..-...- ( -20.50) >DroSec_CAF1 26970 111 + 1 ACAGCGUGAAAUGU-U-----UUUUGUUUUGACUCGCUU-UUCCCGGCGAGUGAGCUCCACUUUUCCACACAAUUCUCACAAUUAAAAACAUAAGAAGAAAACGGAA-AACGCAC-ACAA .....(((...(((-(-----(((((((((...(((((.-.....)))))(((((....................)))))..................)))))))))-))))..)-)).. ( -23.05) >DroYak_CAF1 26867 117 + 1 ACAGUGUCAAAUGU-UUCUUUUUUUGUUUUGACUCGCUU-UUCCCGGCUAGUGAGCUCCACUUUUCCACACAAUUCUCACAAUUAAAAACAUAAGAAGAAAACAGAA-AACGCAUGAAAA .((((((....(((-((((((((.((((((.....(((.-.....)))..(((((....................))))).....)))))).)))))).)))))...-.)))).)).... ( -23.05) >DroAna_CAF1 25191 110 + 1 ACAGCGUAAAAUGU-U-----UUUUGUUUUGUUUCGCUUUUUCCCAGUGAGAGAGCUCCACUUUUCCAAAUAGUUCUCACAAUUAAAAAUAAAAGAAGAAAACAGAA-AACGCAC-UC-- ...((((....(((-(-----((((.((((((((((((.......)))))(((((((..............))))))).........)))))))..))))))))...-.))))..-..-- ( -25.44) >DroPer_CAF1 42959 116 + 1 ACAGCGUCAAAUGU-UUUUAUUUUUUUUUUUGUUUUCUU-UGGCCAGAGAGCUAGCUCCACUUUUCCAUACAAUUCUCACAAUUAUAUUUAUAAGAAGAAAACGGAAAAACGCAC-ACA- ...((((.(((((.-...)))))..((((((((((((((-(((..((.(((....)))..))...))...............((((....)))))))))))))))))))))))..-...- ( -20.50) >consensus ACAGCGUCAAAUGU_U_____UUUUGUUUUGACUCGCUU_UUCCCAGCGAGUGAGCUCCACUUUUCCACACAAUUCUCACAAUUAAAAACAUAAGAAGAAAACAGAA_AACGCAC_ACA_ ...((((..............(((((((((.(((((((.......))))))).....................((((................)))).)))))))))..))))....... (-13.72 = -12.28 + -1.44)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:30:05 2006