
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 6,983,286 – 6,983,435 |
| Length | 149 |
| Max. P | 0.781768 |

| Location | 6,983,286 – 6,983,395 |
|---|---|
| Length | 109 |
| Sequences | 6 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 81.58 |
| Mean single sequence MFE | -36.55 |
| Consensus MFE | -29.50 |
| Energy contribution | -29.42 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.752463 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 6983286 109 - 22407834 UGCCACCGGCAAGACGCGACUGGUUACCGAAUGUCUGCGCGGUGGCAACUGGAGUGUCUCCUACAUACCCAAGUGUCAGGGUGAGUG-UGCCUACCUA-UUCCAU--AAU-UGG- (((((((((((.((((....(((...)))..))))))).))))))))..(((((((.....(((((((((........))))..)))-))......))-))))).--...-...- ( -37.10) >DroVir_CAF1 9677 110 - 1 UGCCACCGGCAAGACGCGUCUGGUUACGGAAUGUUUGCCCGGUGGCAACUGGAGUGUCUCCUACAUACCAAAGUGUCAGGGUAAGUCG---UUAGUUUCAUUCAUUCCAA-UGC- (((((((((((((.....((((....))))...)))).)))))))))..(((((((..((((((((......)))).))))......(---.......)...))))))).-...- ( -37.20) >DroGri_CAF1 10076 110 - 1 UGCCACCGGCAAGACGCGUCUGGUAACUGAAUGUUUGCCCGGUGGCAACUGGAGUGUCUCCUACAUACCAAAGUGUCAGGGUGAGUC-CGCAAAACACAAUUCAU--AUC-UGA- (((((((((((((.....((........))...)))).)))))))))..(((.((((.....)))).)))..((((....(((....-)))...)))).......--...-...- ( -32.60) >DroWil_CAF1 9589 108 - 1 UGCCACCGGCAAGACGCGUCUGGUAACGGAAUGUUUGCCUGGUGGCAACUGGAGUGUCUCCUACAUACCAAAGUGUCAGGGUAAGUA-AAAAAAUAAA---UCAG--AUUAUAU- (((((((((((((.....((((....))))...)))))).))))))).((((..(((....(((.((((..........)))).)))-.....)))..---))))--.......- ( -33.80) >DroMoj_CAF1 9868 113 - 1 UGCCACCGGCAAGACGCGUCUGGUUACGGAAUGUUUGCCUGGUGGCAACUGGAGUGUCUCCUACAUACCAAAGUGUCAGGGUAAGUCGCGCUUAGCUCAGUUCAAUUAAG-UGA- .((((((((((((.....((((....))))...)))))).))))))(((((.(((...((((((((......)))).))))((((.....))))))))))))........-...- ( -38.20) >DroAna_CAF1 9611 108 - 1 UGCCACCGGCAAGACGCGUCUGGUUACGGAAUGUUUGCGCGGUGGCAACUGGAGUGUCUCCUACAUACCGAAGUGUCAGGGUAAGUC-UCAAUUGAU---UUAUU--UUC-UGGC (((((((((((((.....((((....))))...))))).))))))))..(((.((((.....)))).)))....(((((((((((((-......)))---)))).--.))-)))) ( -40.40) >consensus UGCCACCGGCAAGACGCGUCUGGUUACGGAAUGUUUGCCCGGUGGCAACUGGAGUGUCUCCUACAUACCAAAGUGUCAGGGUAAGUC_CGCAUAGCUA__UUCAU__AAC_UGA_ (((((((((((.((((..((((....)))).))))))).))))))))...((((...)))).((.((((..........)))).))............................. (-29.50 = -29.42 + -0.08)



| Location | 6,983,315 – 6,983,435 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.67 |
| Mean single sequence MFE | -43.62 |
| Consensus MFE | -39.10 |
| Energy contribution | -38.47 |
| Covariance contribution | -0.64 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.781768 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 6983315 120 - 22407834 GUACGCUGGUCAGCUUUACAUGUCCCAUUGGACAGGAGUUUGCCACCGGCAAGACGCGACUGGUUACCGAAUGUCUGCGCGGUGGCAACUGGAGUGUCUCCUACAUACCCAAGUGUCAGG .....((((...((((....(((((....)))))((((((.((((((((((.((((....(((...)))..))))))).)))))))))))((((...)))).......)))))).)))). ( -40.80) >DroVir_CAF1 9707 120 - 1 GCACGCUGGUCAGCUUUACAUGUCCCAUUGGACAGGAGUUUGCCACCGGCAAGACGCGUCUGGUUACGGAAUGUUUGCCCGGUGGCAACUGGAGUGUCUCCUACAUACCAAAGUGUCAGG ((((.(..((.(((((....(((((....))))).)))))(((((((((((((.....((((....))))...)))).)))))))))))..).))))..(((((((......)))).))) ( -46.00) >DroGri_CAF1 10106 120 - 1 GCACGCUGGUCAGCUUCACAUGUCCCAUUGGACAGGAGUUUGCCACCGGCAAGACGCGUCUGGUAACUGAAUGUUUGCCCGGUGGCAACUGGAGUGUCUCCUACAUACCAAAGUGUCAGG ((((.(..((.((((((...(((((....)))))))))))(((((((((((((.....((........))...)))).)))))))))))..).))))..(((((((......)))).))) ( -43.50) >DroSim_CAF1 10439 120 - 1 GUACGCUGGUCAGCUUUACAUGUCCCAUUGGACAGGAGUUUGCCACCGGCAAGACGCGACUGGUUACUGAAUGUCUGCGCGGUGGCAACUGGAGUGUCUCCUACAUACCCAAGUGUCAGG .....((((...((((....(((((....)))))((((((.((((((((((.((((.....(....)....))))))).)))))))))))((((...)))).......)))))).)))). ( -40.10) >DroWil_CAF1 9617 120 - 1 GUACGCUGGUCAGCUUUACAUGUCCCAUUGGACAGGAGUUUGCCACCGGCAAGACGCGUCUGGUAACGGAAUGUUUGCCUGGUGGCAACUGGAGUGUCUCCUACAUACCAAAGUGUCAGG .....((((...(((((...(((((....)))))((((((.((((((((((((.....((((....))))...)))))).))))))))))((((...))))......))))))).)))). ( -45.30) >DroMoj_CAF1 9901 120 - 1 GCACGCUGGUCAGCUUUACAUGUCCCAUUGGACAAGAGUUUGCCACCGGCAAGACGCGUCUGGUUACGGAAUGUUUGCCUGGUGGCAACUGGAGUGUCUCCUACAUACCAAAGUGUCAGG ((((.(..((.((((((...(((((....)))))))))))(((((((((((((.....((((....))))...)))))).)))))))))..).))))..(((((((......)))).))) ( -46.00) >consensus GCACGCUGGUCAGCUUUACAUGUCCCAUUGGACAGGAGUUUGCCACCGGCAAGACGCGUCUGGUUACGGAAUGUUUGCCCGGUGGCAACUGGAGUGUCUCCUACAUACCAAAGUGUCAGG .(((.(..((.((((((...(((((....)))))))))))(((((((((((.(((...((.(....).))..)))))).))))))))))..).)))...(((((((......)))).))) (-39.10 = -38.47 + -0.64)

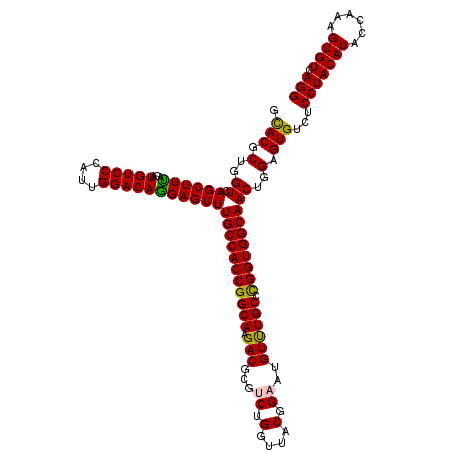

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:29:59 2006