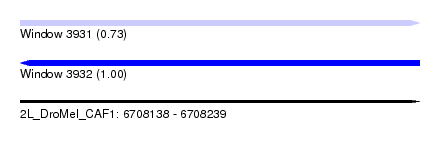

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 6,708,138 – 6,708,239 |
| Length | 101 |
| Max. P | 0.999382 |

| Location | 6,708,138 – 6,708,239 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 91.70 |
| Mean single sequence MFE | -32.46 |
| Consensus MFE | -24.92 |
| Energy contribution | -26.72 |
| Covariance contribution | 1.80 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.727602 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 6708138 101 + 22407834 CUUCCAGGCACUUGAAUAUGGAAUGCACUUCGCACCUGGCCACUUGGCGCACUCAUUUUCCGAGUGAGCGGAGUGUAGCCCAGAUUUUGGCCACACGAA-UA ......(((....((((.(((..((((((((((.....(((....))).(((((.......))))).))))))))))..))).))))..))).......-.. ( -39.20) >DroSec_CAF1 20805 101 + 1 CUUCCACGCACUUGAAUAUGGAAUGCACUUCGCAACUGGCCACUUGGCGCACUGAUUUUCCGAGUGAGCGGAGUGUAGCCCAUAUUUUGGCCACACGAU-UG .......((....((((((((..((((((((((.....(((....))).(((((......).)))).))))))))))..))))))))..))........-.. ( -35.60) >DroSim_CAF1 20615 101 + 1 CUUCCACGCACUUGAAUAUGGAAUGCACUUCGCACCUGGUCACUUGGCGCACUCAUUUUCCGAGUGAGCGGAGUGUAGCCCAGAUUUUGGCCACACGAA-UG .......((....((((.(((..((((((((((.....(((....))).(((((.......))))).))))))))))..))).))))..))........-.. ( -32.40) >DroEre_CAF1 21877 99 + 1 CUUCCACGCACUUGAAUAUGGAAUGCACUUCGCACCUGGCCACUUUGCGCAUUCAUUUUCCGAGUGAGCGGA--AUAGCCCAGAUUGUGGCCACACGAU-CC .((((.(.((((((......((((((.(...((.....))......).))))))......)))))).).)))--).......(((((((....))))))-). ( -29.40) >DroYak_CAF1 21770 100 + 1 CUUCCACGCACUUGAAUACGGAAUGCACUUCGCACCUGGCCACUUUGCGCAUUCAUUUUCCGAGUGAGCGGC--GUAGCCCAGAUUUUGGCCACACGAAGCA .......(((.((.......)).))).(((((....((((((....((.(((((.......))))).))(((--...))).......))))))..))))).. ( -25.70) >consensus CUUCCACGCACUUGAAUAUGGAAUGCACUUCGCACCUGGCCACUUGGCGCACUCAUUUUCCGAGUGAGCGGAGUGUAGCCCAGAUUUUGGCCACACGAA_UA .......((....((((.(((..((((((((((.....(((....))).(((((.......))))).))))))))))..))).))))..))........... (-24.92 = -26.72 + 1.80)

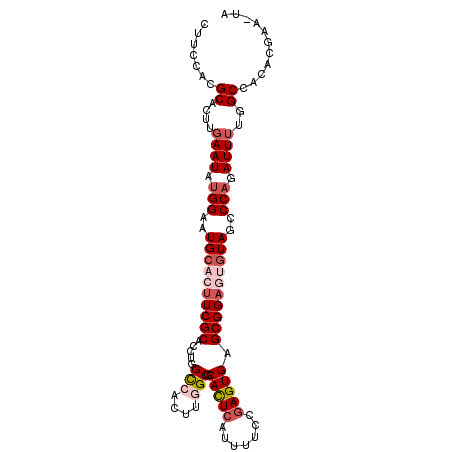
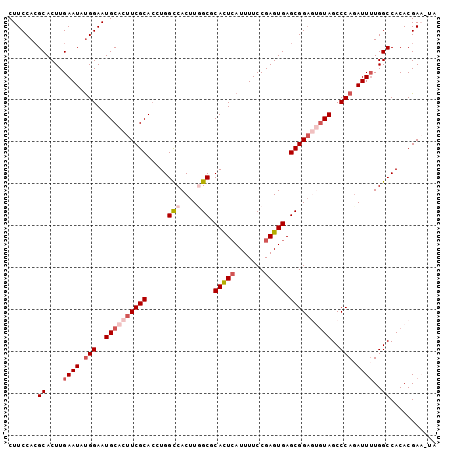
| Location | 6,708,138 – 6,708,239 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 91.70 |
| Mean single sequence MFE | -36.76 |
| Consensus MFE | -33.46 |
| Energy contribution | -34.46 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.31 |
| Structure conservation index | 0.91 |
| SVM decision value | 3.56 |
| SVM RNA-class probability | 0.999382 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 6708138 101 - 22407834 UA-UUCGUGUGGCCAAAAUCUGGGCUACACUCCGCUCACUCGGAAAAUGAGUGCGCCAAGUGGCCAGGUGCGAAGUGCAUUCCAUAUUCAAGUGCCUGGAAG ..-...((((((((........)))))))).(((((((((((.....)))))).....)))))(((((..(((((((.....))).)))..)..)))))... ( -37.30) >DroSec_CAF1 20805 101 - 1 CA-AUCGUGUGGCCAAAAUAUGGGCUACACUCCGCUCACUCGGAAAAUCAGUGCGCCAAGUGGCCAGUUGCGAAGUGCAUUCCAUAUUCAAGUGCGUGGAAG ..-...((((((((........))))))))(((((.((((.(((.....(((((((...(..(....)..)...))))))).....))).)))).))))).. ( -38.10) >DroSim_CAF1 20615 101 - 1 CA-UUCGUGUGGCCAAAAUCUGGGCUACACUCCGCUCACUCGGAAAAUGAGUGCGCCAAGUGACCAGGUGCGAAGUGCAUUCCAUAUUCAAGUGCGUGGAAG ..-...((((((((........))))))))(((((.((((.(((....((((((((...((........))...))))))))....))).)))).))))).. ( -39.20) >DroEre_CAF1 21877 99 - 1 GG-AUCGUGUGGCCACAAUCUGGGCUAU--UCCGCUCACUCGGAAAAUGAAUGCGCAAAGUGGCCAGGUGCGAAGUGCAUUCCAUAUUCAAGUGCGUGGAAG ..-......(((((........)))))(--(((((.((((.(((....((((((((...((........))...))))))))....))).)))).)))))). ( -32.60) >DroYak_CAF1 21770 100 - 1 UGCUUCGUGUGGCCAAAAUCUGGGCUAC--GCCGCUCACUCGGAAAAUGAAUGCGCAAAGUGGCCAGGUGCGAAGUGCAUUCCGUAUUCAAGUGCGUGGAAG ..((((((((((((........))))))--))(((.((((.(((....((((((((...((........))...))))))))....))).)))).))))))) ( -36.60) >consensus CA_UUCGUGUGGCCAAAAUCUGGGCUACACUCCGCUCACUCGGAAAAUGAGUGCGCCAAGUGGCCAGGUGCGAAGUGCAUUCCAUAUUCAAGUGCGUGGAAG ......((((((((........))))))))(((((.((((.(((....((((((((...((........))...))))))))....))).)))).))))).. (-33.46 = -34.46 + 1.00)

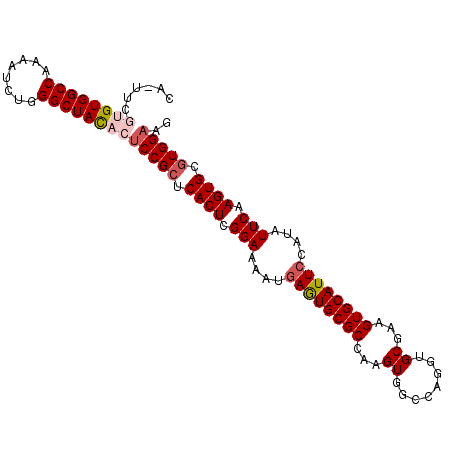
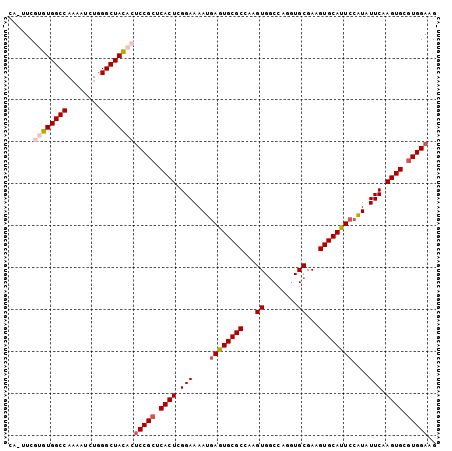
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:28:14 2006