
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 6,698,399 – 6,698,495 |
| Length | 96 |
| Max. P | 0.990175 |

| Location | 6,698,399 – 6,698,495 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 87.64 |
| Mean single sequence MFE | -30.16 |
| Consensus MFE | -21.76 |
| Energy contribution | -21.88 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.58 |
| Structure conservation index | 0.72 |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.929329 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
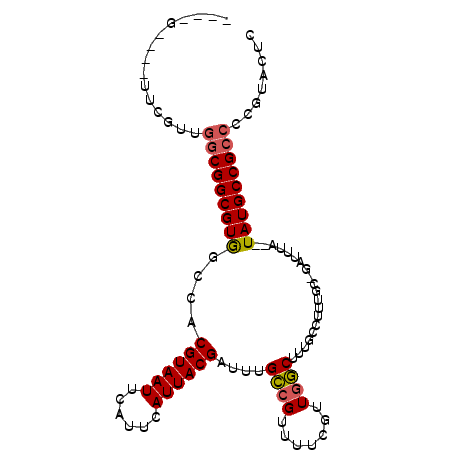
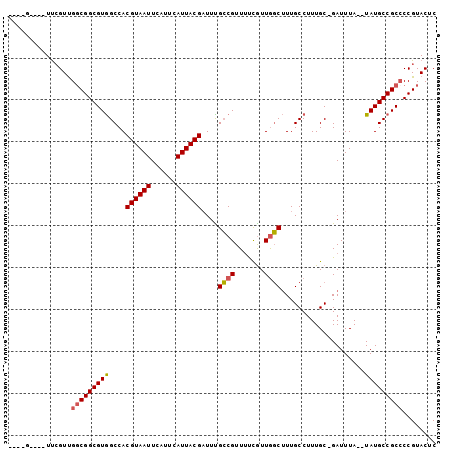
>2L_DroMel_CAF1 6698399 96 + 22407834 ----G--CGUUCGUUGGCGGCGUGGCCACGUAAUUCAUUCAUUACGAUUUGCCGUUUUCGUUGGCUUUGCCUUUGCCGAUUUAUAUAUGCCGCCCCGUACUC ----.--.((.((..(((((((((((..((((((......))))))....)))......((((((.........))))))......)))))))).)).)).. ( -31.50) >DroSec_CAF1 11403 92 + 1 ----G----UUCGUUGGCGGCGUGGCCACGUAAUUCAUUCAUUACGAUUUGCCGUUUUUGUUGGCUUUGCCUUUGCCGAUUUA--UAUGCCGCACCGUACUC ----(----(.((...((((((((((..((((((......))))))....)))......((((((.........))))))...--.)))))))..)).)).. ( -28.80) >DroEre_CAF1 11568 91 + 1 ----G----UUCGUUGGCGGCGUGGCCACGUAAUUCAUUCAUUACGAUUCGCCGUUUUCGUUGGCUUUGCCUUUGC-GAUUUA--UAUGCCGCCCCGUACUC ----(----(.((..(((((((((((..((((((......))))))....)))....((((.(((...)))...))-))....--.)))))))).)).)).. ( -31.50) >DroYak_CAF1 11663 91 + 1 ----G----UUCGUUGGCGGCGUGGCCACGUAAUUCAUUCAUUACGAUUCGCCGUUUUCGUUGGCUUUGCCUUUGC-GAUUUA--UAUGCCGCCCCGUACUC ----(----(.((..(((((((((((..((((((......))))))....)))....((((.(((...)))...))-))....--.)))))))).)).)).. ( -31.50) >DroAna_CAF1 15799 99 + 1 UCUGGUAGGUCGGUCGUCGGCGUAGCCACGUAAUUCAUUCAUUACGUUUUGGCGUUGCUAUUACCAUUGCCUUUGC-GAUUUA--UAUGCCGCCCCGUACUC .......((((((..(.(((((((((((((((((......)))))))...)))(((((................))-)))...--)))))))).))).))). ( -27.49) >consensus ____G____UUCGUUGGCGGCGUGGCCACGUAAUUCAUUCAUUACGAUUUGCCGUUUUCGUUGGCUUUGCCUUUGC_GAUUUA__UAUGCCGCCCCGUACUC ...............(((((((((....((((((......))))))....((((.......))))....................)))))))))........ (-21.76 = -21.88 + 0.12)

| Location | 6,698,399 – 6,698,495 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 87.64 |
| Mean single sequence MFE | -28.82 |
| Consensus MFE | -22.44 |
| Energy contribution | -22.92 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.82 |
| Structure conservation index | 0.78 |
| SVM decision value | 2.20 |
| SVM RNA-class probability | 0.990175 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
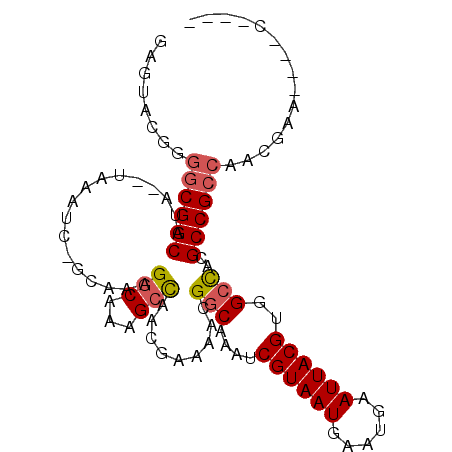
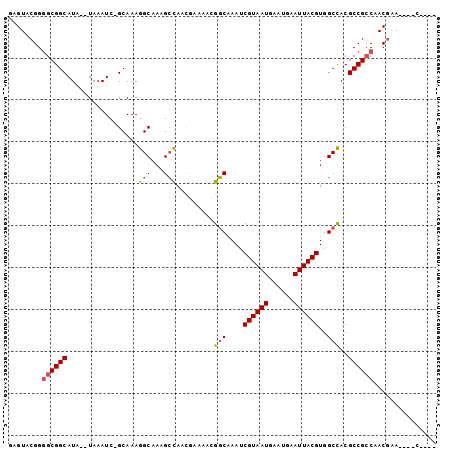
>2L_DroMel_CAF1 6698399 96 - 22407834 GAGUACGGGGCGGCAUAUAUAAAUCGGCAAAGGCAAAGCCAACGAAAACGGCAAAUCGUAAUGAAUGAAUUACGUGGCCACGCCGCCAACGAACG--C---- ..((.((.((((((...........(((.........))).........(((....((((((......))))))..)))..))))))..)).)).--.---- ( -32.10) >DroSec_CAF1 11403 92 - 1 GAGUACGGUGCGGCAUA--UAAAUCGGCAAAGGCAAAGCCAACAAAAACGGCAAAUCGUAAUGAAUGAAUUACGUGGCCACGCCGCCAACGAA----C---- .....((..(((((...--......(((.........))).........(((....((((((......))))))..)))..)))))...))..----.---- ( -27.90) >DroEre_CAF1 11568 91 - 1 GAGUACGGGGCGGCAUA--UAAAUC-GCAAAGGCAAAGCCAACGAAAACGGCGAAUCGUAAUGAAUGAAUUACGUGGCCACGCCGCCAACGAA----C---- .....((.((((((...--....((-(....(((...)))..)))....(((.(..((((((......))))))).)))..))))))..))..----.---- ( -29.20) >DroYak_CAF1 11663 91 - 1 GAGUACGGGGCGGCAUA--UAAAUC-GCAAAGGCAAAGCCAACGAAAACGGCGAAUCGUAAUGAAUGAAUUACGUGGCCACGCCGCCAACGAA----C---- .....((.((((((...--....((-(....(((...)))..)))....(((.(..((((((......))))))).)))..))))))..))..----.---- ( -29.20) >DroAna_CAF1 15799 99 - 1 GAGUACGGGGCGGCAUA--UAAAUC-GCAAAGGCAAUGGUAAUAGCAACGCCAAAACGUAAUGAAUGAAUUACGUGGCUACGCCGACGACCGACCUACCAGA .((..(((..((((...--......-((....((....))....))...(((...(((((((......))))))))))...))))....)))..))...... ( -25.70) >consensus GAGUACGGGGCGGCAUA__UAAAUC_GCAAAGGCAAAGCCAACGAAAACGGCAAAUCGUAAUGAAUGAAUUACGUGGCCACGCCGCCAACGAA____C____ ........((((((.................(((...))).........(((....((((((......))))))..)))..))))))............... (-22.44 = -22.92 + 0.48)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:28:09 2006