
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 6,525,242 – 6,525,345 |
| Length | 103 |
| Max. P | 0.593453 |

| Location | 6,525,242 – 6,525,345 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 75.91 |
| Mean single sequence MFE | -23.87 |
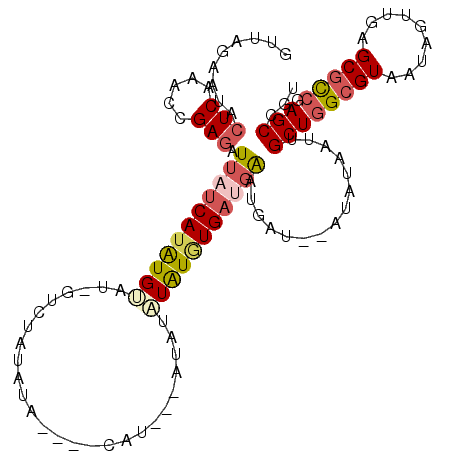
| Consensus MFE | -13.32 |
| Energy contribution | -14.24 |
| Covariance contribution | 0.92 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.56 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.593453 |
| Prediction | RNA |
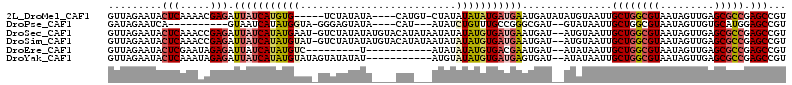
Download alignment: ClustalW | MAF
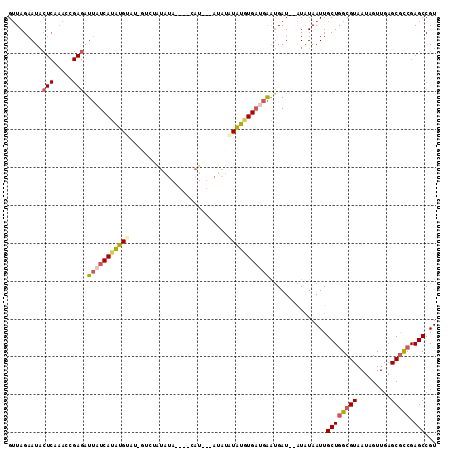
>2L_DroMel_CAF1 6525242 103 - 22407834 GUUAGAAUACUCAAAACGAGAUUAUCAUGUG-----UCUAUAUA----CAUGU-CUAUAUAUAUGAUGAAUGAUAUAUGUAAUUGCUGGCGUAAUAGUUGAGCGCCGAGCCGU ...............(((.(((((.((((((-----((..(((.----(((((-......))))))))...)))))))))))))((((((((.........))))).)))))) ( -24.00) >DroPse_CAF1 18835 93 - 1 GAUAGAAUCA----------GUAAUCAUAUGGUA-GGGAGUAUA----CAU---AUAUCUGUUUGCCGGGCGAU--GUAUAAUUGCUGGCGUAAUAGUUGUGCAUGGAGCCGU ..........----------.........(((((-((.(((((.----...---))).)).)))))))(((.((--(((((((((.........)))))))))))...))).. ( -20.90) >DroSec_CAF1 12335 110 - 1 GUUAGAAUACUCAAACCGAGAUUAUCAUAUGAAU-GUCUAUAUAUGUACAUAUAAUAUAUAUGUGAUGAAUGAU--AUGUAAUUGCUGGCGUAAUAGUUGAGCGCCGAGCCGU .........(((.....))).((((((((((.((-((.(((((......))))))))).)))))))))).....--........((((((((.........))))).)))... ( -23.90) >DroSim_CAF1 12223 110 - 1 GUUAGAAUACUCAAACCGAGAUUAUCAUAUGUAU-GUCUAUAUAUGUACAUAUAAUAUAUAUGUGAUGAAUGAU--AUGUAAUUGCUGGCGUAAUAGUUGAGCGCCGAGCCGU .........(((.....))).(((((((((((((-((.(((((......)))))))))))))))))))).....--........((((((((.........))))).)))... ( -28.30) >DroEre_CAF1 9773 91 - 1 GUUAGAAUACUCGAAUAGAGAUUAUCAUAUGUC---------U-----------AUAUAUAUGUGACGAAUGAU--AUAUAAUUGCUGGCGUAAUAGUUGAGCGCCGAGCCGU .........(((.....)))..((((((.((((---------.-----------..........)))).)))))--).......((((((((.........))))).)))... ( -21.00) >DroYak_CAF1 11479 100 - 1 GUUAGAAUACUCAAAUAGAGAUUAUCAUAUGUAUAGUAUAUAU-----------AUGUAUAUGUGAUGAGUGAU--AUAUAAUUGCUGGCGUAAUAGUUGAGCGCCGAGCCGU .........(((.....))).((((((((((((((........-----------.)))))))))))))).....--........((((((((.........))))).)))... ( -25.10) >consensus GUUAGAAUACUCAAACCGAGAUUAUCAUAUGUAU_GUCUAUAUA____CAU___AUAUAUAUGUGAUGAAUGAU__AUAUAAUUGCUGGCGUAAUAGUUGAGCGCCGAGCCGU .........(((.....))).(((((((((((..........................)))))))))))...............((((((((.........))))).)))... (-13.32 = -14.24 + 0.92)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:27:02 2006