
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 6,423,248 – 6,423,346 |
| Length | 98 |
| Max. P | 0.923688 |

| Location | 6,423,248 – 6,423,346 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 77.74 |
| Mean single sequence MFE | -33.22 |
| Consensus MFE | -21.88 |
| Energy contribution | -22.97 |
| Covariance contribution | 1.09 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.798889 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
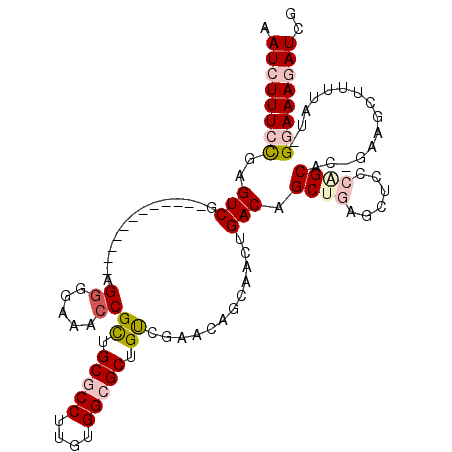
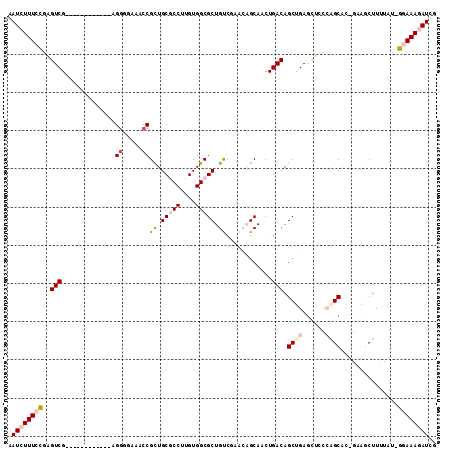
>2L_DroMel_CAF1 6423248 98 + 22407834 AAUCUUUCCGAGUCG------------AGAUGAAACCGCUGCGCCCUGUGGCGCUGUCGAACAGCAACUGACAGCUGAAUUCCCAGCAC-GAAGCUUUUAU-GGAAAGAUCG .(((((((((.(((.------------((.((....(((.(((((....))))).).)).....)).))))).((((......))))..-..........)-)))))))).. ( -31.40) >DroPse_CAF1 19094 106 + 1 UAUGUUU-UGCGUCGUUUUUUCGAAAUAGGAACAACCGUUGCACCUCGUGGCGCUGCC-----GCAACUGACAGCCGAGCGGCAAGCAUUGAAUCUUUUAUUGGAAAAAUCU .......-......((((((((((...((((......((..(.....)..))((((((-----((.............))))).)))......))))...)))))))))).. ( -26.12) >DroSim_CAF1 9762 98 + 1 AAUCUUUCCGAGUCG------------AGGGGAAACCGCUGCGCCUUGUGGCGCUGUCGAACAGCAACUGACAGCUGAGUUCCCAGCAC-GGAGCUUUUAU-GGAAAGAUCG .(((((((((..(((------------(.((....))...(((((....)))))..)))).((((........))))((((((......-))))))....)-)))))))).. ( -40.00) >DroEre_CAF1 10542 98 + 1 AAUCUUUCCGAGUCG------------UGGGGAAACCGCUGCGCCUUGUGGCGCUGUCGAACAGCAACUGACAGCUGAGUUCCCAGCAC-GAAGCUUUUAU-GGAAAGAUCG .((((((((((((((------------(((((((..(((((((((....)))((((.....))))....).)))).)..)))))..)))-)..))))....-)))))))).. ( -38.60) >DroYak_CAF1 10100 98 + 1 AAUCUUUCCGAGUCG------------AGGGGAAACCGCUGCGCCUUGUGGCGCUGUCGAACAGCAACUGACAGCUGAGCUCCCGGCAC-GAAGCUUUUAU-GGAAAGAUCG .(((((((((.....------------..((....))((((((((....)))))..(((..((((........)))).((.....)).)-))))).....)-)))))))).. ( -37.10) >DroPer_CAF1 19466 106 + 1 UAUGUUU-UGCGUCGUUUUUUCGAAAUAGGAACAACCGUUGCACCUCGUGGCGCUGCC-----GCAACUGACAGCCGAGCGGCAAGCAUUGAAUCUUUUAUUGGAAAAAUCU .......-......((((((((((...((((......((..(.....)..))((((((-----((.............))))).)))......))))...)))))))))).. ( -26.12) >consensus AAUCUUUCCGAGUCG____________AGGGGAAACCGCUGCGCCUUGUGGCGCUGUCGAACAGCAACUGACAGCUGAGCUCCCAGCAC_GAAGCUUUUAU_GGAAAGAUCG .((((((((..(((..............((.....))((.(((((....))))).))............))).((((......))))...............)))))))).. (-21.88 = -22.97 + 1.09)

| Location | 6,423,248 – 6,423,346 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 77.74 |
| Mean single sequence MFE | -33.62 |
| Consensus MFE | -21.48 |
| Energy contribution | -23.03 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.64 |
| SVM decision value | 1.16 |
| SVM RNA-class probability | 0.923688 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
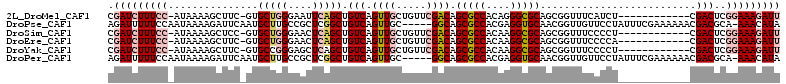
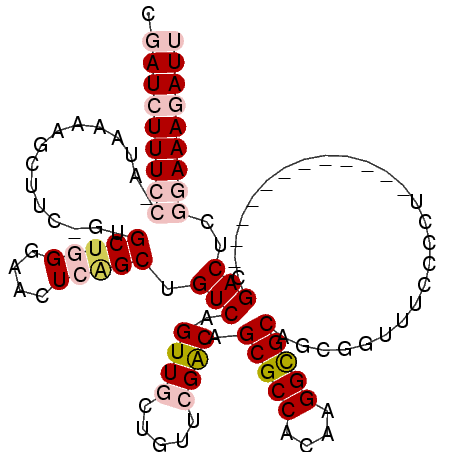
>2L_DroMel_CAF1 6423248 98 - 22407834 CGAUCUUUCC-AUAAAAGCUUC-GUGCUGGGAAUUCAGCUGUCAGUUGCUGUUCGACAGCGCCACAGGGCGCAGCGGUUUCAUCU------------CGACUCGGAAAGAUU .(((((((((-......(((.(-((.(((((......((((((...........)))))).)).))).))).)))((((......------------.)))).))))))))) ( -35.10) >DroPse_CAF1 19094 106 - 1 AGAUUUUUCCAAUAAAAGAUUCAAUGCUUGCCGCUCGGCUGUCAGUUGC-----GGCAGCGCCACGAGGUGCAACGGUUGUUCCUAUUUCGAAAAAACGACGCA-AAACAUA .(((((((......)))))))...(((.((((((...((.....)).))-----))))(((((....)))))....((((((.............)))))))))-....... ( -25.92) >DroSim_CAF1 9762 98 - 1 CGAUCUUUCC-AUAAAAGCUCC-GUGCUGGGAACUCAGCUGUCAGUUGCUGUUCGACAGCGCCACAAGGCGCAGCGGUUUCCCCU------------CGACUCGGAAAGAUU .(((((((((-.....(((...-..)))(((((..(.(((((.....((((.....))))(((....)))))))))..)))))..------------......))))))))) ( -38.80) >DroEre_CAF1 10542 98 - 1 CGAUCUUUCC-AUAAAAGCUUC-GUGCUGGGAACUCAGCUGUCAGUUGCUGUUCGACAGCGCCACAAGGCGCAGCGGUUUCCCCA------------CGACUCGGAAAGAUU .(((((((((-.....((..((-(((..(((((((..(((((.....((((.....))))(((....))))))))))))))).))------------))))).))))))))) ( -38.80) >DroYak_CAF1 10100 98 - 1 CGAUCUUUCC-AUAAAAGCUUC-GUGCCGGGAGCUCAGCUGUCAGUUGCUGUUCGACAGCGCCACAAGGCGCAGCGGUUUCCCCU------------CGACUCGGAAAGAUU .(((((((((-......((...-..)).(((((..(.(((((.....((((.....))))(((....)))))))))..)))))..------------......))))))))) ( -37.20) >DroPer_CAF1 19466 106 - 1 AGAUUUUUCCAAUAAAAGAUUCAAUGCUUGCCGCUCGGCUGUCAGUUGC-----GGCAGCGCCACGAGGUGCAACGGUUGUUCCUAUUUCGAAAAAACGACGCA-AAACAUA .(((((((......)))))))...(((.((((((...((.....)).))-----))))(((((....)))))....((((((.............)))))))))-....... ( -25.92) >consensus CGAUCUUUCC_AUAAAAGCUUC_GUGCUGGGAACUCAGCUGUCAGUUGCUGUUCGACAGCGCCACAAGGCGCAGCGGUUUCCCCU____________CGACUCGGAAAGAUU .(((((((((...............(((((....))))).(((.((((.....)))).(((((....)))))..........................)))..))))))))) (-21.48 = -23.03 + 1.56)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:26:27 2006