
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 6,334,809 – 6,334,916 |
| Length | 107 |
| Max. P | 0.772616 |

| Location | 6,334,809 – 6,334,916 |
|---|---|
| Length | 107 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 76.93 |
| Mean single sequence MFE | -45.58 |
| Consensus MFE | -26.59 |
| Energy contribution | -26.87 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.772616 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
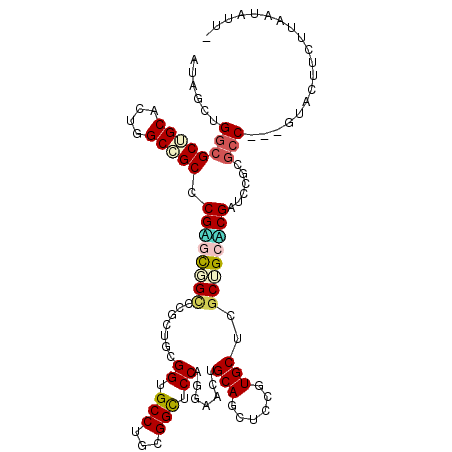
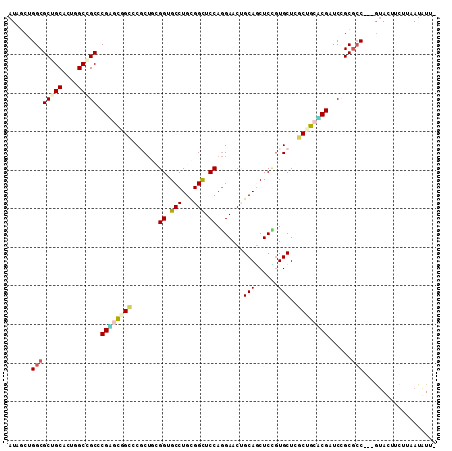
>2L_DroMel_CAF1 6334809 107 + 22407834 AUAUCUGGCGCUGCACUGGCGGCUCGAUUGGUCCGCUUAGGAGCCUGCGGUUCCAGGAACAGCAGCUCCGUGCUUGCUGCUCGGUCUGCGCC---GUACUUCUUUAUGUU- ......(((((.(.(((((((((..((....)).((...(((((.(((.((((...)))).))))))))..))..))))).))))).)))))---...............- ( -41.80) >DroVir_CAF1 44902 108 + 1 AUUGCGGGCGCUGCGCUGGCGGCGCGUGCCGCACGCUGCGGUGCCUGUGGCUCCAGGAACUGCAGCUCCGUGCUGGCCGCACGAUCGGCGCC---GUAUUUCUUAAUAUUU ..((((.(((((((....)))))))((((((...((((((((.((((......)))).))))))))..(((((.....)))))..)))))))---)))............. ( -54.50) >DroPse_CAF1 64843 108 + 1 AUGGCCGGCGCAGCACUGGCCGCCCGGGCCGCCCUCUGCGGUGCCUGCGGCUCCAGGAACUGCAUCUCGCUGCUGACCGCCCGAUCCGCUGC---GAACUUUUGGAUAUUC .(((.(((((((((.((((..((((((((.(((......)))))))).))).))))((........))))))).).))).)))(((((....---.......))))).... ( -41.00) >DroGri_CAF1 19550 108 + 1 AUUGCUGGCGCGGCACUUGCCGCACGUGCGGCACGCAACGGUGCCUGUGGCUCCAAGAACUGCAACUCUGUGCUCGCUGCACGAUCCGCGCC---GUAUUUCUUAAUAUUU ..(((.((((((((....)))...((((((((..(((..((.(((...))).)).(((........))).)))..))))))))....)))))---)))............. ( -42.00) >DroSim_CAF1 27864 107 + 1 AUAUCUGGCGCUGCACUGGCGGCUCGGUUGGUCCGCUUAGGAGCCUGCGGUUCCAGGAACAGCAGCUCCGUGCUUGCUGCUCGGUCUGCGCC---GUACUUUUUUAUGUU- ......(((((.(.(((((((((..((.....))((...(((((.(((.((((...)))).))))))))..))..))))).))))).)))))---...............- ( -44.50) >DroMoj_CAF1 53315 111 + 1 AUGGCGGGCGCUGCGCUGGCCGCCCGAGCUGCUCUCUGCGGCGCCUGUGGCUCCAGGAACUGCAGCUCUGUGCUCGCAGCACGAUCCGCGCCCGCAUAUUUCUUAAUAUUC ...((((((((((.((((...(((.(((((((..(((..((.(((...))).)).)))...))))))).).))...)))).))....))))))))................ ( -49.70) >consensus AUAGCUGGCGCUGCACUGGCCGCCCGAGCGGCCCGCUGCGGUGCCUGCGGCUCCAGGAACUGCAGCUCCGUGCUCGCUGCACGAUCCGCGCC___GUACUUCUUAAUAUU_ ......((((((((....))))).((((((((.......((.(((...))).)).......(((......)))..))))))))......)))................... (-26.59 = -26.87 + 0.28)

| Location | 6,334,809 – 6,334,916 |
|---|---|
| Length | 107 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 76.93 |
| Mean single sequence MFE | -44.20 |
| Consensus MFE | -24.08 |
| Energy contribution | -24.88 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.35 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.54 |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.748085 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
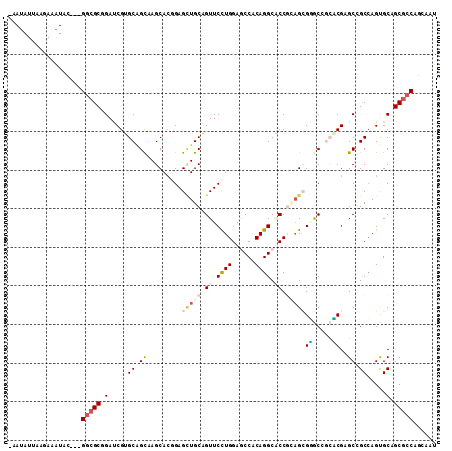
>2L_DroMel_CAF1 6334809 107 - 22407834 -AACAUAAAGAAGUAC---GGCGCAGACCGAGCAGCAAGCACGGAGCUGCUGUUCCUGGAACCGCAGGCUCCUAAGCGGACCAAUCGAGCCGCCAGUGCAGCGCCAGAUAU -...............---(((((.(......).....((((((((((((.((((...)))).))).)))))...((((..(....)..))))..)))).)))))...... ( -38.60) >DroVir_CAF1 44902 108 - 1 AAAUAUUAAGAAAUAC---GGCGCCGAUCGUGCGGCCAGCACGGAGCUGCAGUUCCUGGAGCCACAGGCACCGCAGCGUGCGGCACGCGCCGCCAGCGCAGCGCCCGCAAU ................---(((((.....((((((((.((((...(((((.((.((((......)))).)).)))))))))))).))))).((....)).)))))...... ( -47.00) >DroPse_CAF1 64843 108 - 1 GAAUAUCCAAAAGUUC---GCAGCGGAUCGGGCGGUCAGCAGCGAGAUGCAGUUCCUGGAGCCGCAGGCACCGCAGAGGGCGGCCCGGGCGGCCAGUGCUGCGCCGGCCAU .....(((.....(((---((.((.((((....)))).)).))))).(((.((.((((......)))).)).)))..))).((((.(((((((....))))).)))))).. ( -44.50) >DroGri_CAF1 19550 108 - 1 AAAUAUUAAGAAAUAC---GGCGCGGAUCGUGCAGCGAGCACAGAGUUGCAGUUCUUGGAGCCACAGGCACCGUUGCGUGCCGCACGUGCGGCAAGUGCCGCGCCAGCAAU ................---(((((((.....((((((.(((......))).((.((((......)))).)))))))).((((((....))))))....)))))))...... ( -43.80) >DroSim_CAF1 27864 107 - 1 -AACAUAAAAAAGUAC---GGCGCAGACCGAGCAGCAAGCACGGAGCUGCUGUUCCUGGAACCGCAGGCUCCUAAGCGGACCAACCGAGCCGCCAGUGCAGCGCCAGAUAU -...............---(((((.(......).....((((((((((((.((((...)))).))).)))))...((((..(....)..))))..)))).)))))...... ( -38.60) >DroMoj_CAF1 53315 111 - 1 GAAUAUUAAGAAAUAUGCGGGCGCGGAUCGUGCUGCGAGCACAGAGCUGCAGUUCCUGGAGCCACAGGCGCCGCAGAGAGCAGCUCGGGCGGCCAGCGCAGCGCCCGCCAU ................((((((((.....((((((.(.((...(((((((..((..(((.(((...))).)))..))..)))))))..))..))))))).))))))))... ( -52.70) >consensus _AAUAUUAAGAAAUAC___GGCGCGGAUCGUGCAGCAAGCACGGAGCUGCAGUUCCUGGAGCCACAGGCACCGCAGCGGGCCGCACGAGCCGCCAGUGCAGCGCCAGCAAU ...................(((((.(.....((.((.........(((((.((.((((......)))).)).)))))((.....))..)).)).....).)))))...... (-24.08 = -24.88 + 0.81)
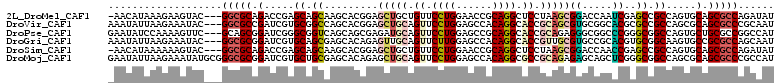


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:25:38 2006