
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 6,302,820 – 6,302,934 |
| Length | 114 |
| Max. P | 0.801117 |

| Location | 6,302,820 – 6,302,934 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 73.94 |
| Mean single sequence MFE | -28.67 |
| Consensus MFE | -19.36 |
| Energy contribution | -19.17 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.801117 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
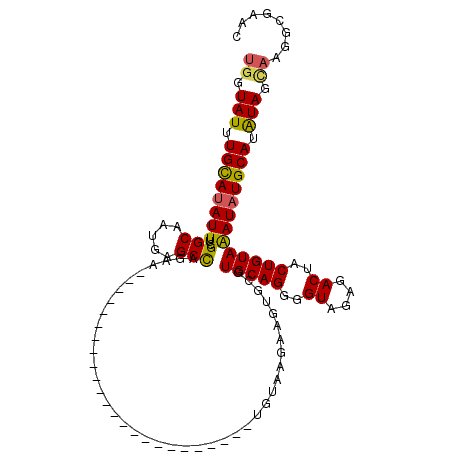
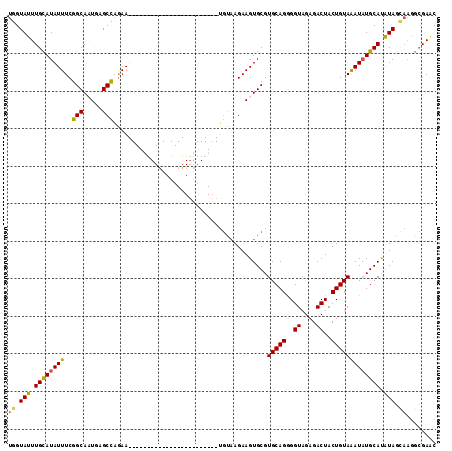
>2L_DroMel_CAF1 6302820 114 + 22407834 UGGUAUUUGUAUAUUUCGGCACUGAGCCAAAAUUGAAUUCUACGCAGGCUGAAUGGUGUAAGAAGUGCGUGCAGGGGUAGAGACUACUGUAAAUAUGCAUAUAGCAAGGCGAAC ((.(((.(((((((((.(((.....))).............(((((..((..........))...)))))((((..((....))..))))))))))))).))).))........ ( -26.10) >DroSec_CAF1 138810 90 + 1 UGGUAUUUGCAUAUUUCGGCAAUGAGCCAGAA------------------------UGUAAGAAGUGCGUGCAGGGGUAGAGACUACUGUAAAUUUGCAUGUAGUAAGGCGAAC .....((((((((((..(((.....)))..))------------------------)))....(.((((((((((..((.((....)).))..)))))))))).)...))))). ( -27.20) >DroSim_CAF1 160591 90 + 1 UGGUAUUUGCAUAUUUCGGCAAUGAGCCAGAA------------------------UGUAAGAAGUGCGUGCAGGGGUAGAGACUACUGUAAAUAUGCAUGUAGCAAGGCGAAC ..((.((((((((((..(((.....)))..))------------------------)))......((((((((....((.((....)).))....))))))))))))))).... ( -25.30) >DroEre_CAF1 138793 110 + 1 UGGUAUUUGCAUAUUUCGGCACUGCGCUCGAAUUGAGCUCCACGCAGGGGGAAUCGGUCCAGAAGUGCGUGCAGAGGUAGAGACUACUGUAGAUAUGCACAUAC----GAGAGC ..((((.((((((((((.((((.((((((..(((((.((((.....))))...)))))...).)))))))))...(((((...)))))).))))))))).))))----...... ( -36.10) >consensus UGGUAUUUGCAUAUUUCGGCAAUGAGCCAGAA________________________UGUAAGAAGUGCGUGCAGGGGUAGAGACUACUGUAAAUAUGCAUAUAGCAAGGCGAAC ((.(((.((((((((..(((.....))).........................................(((((..((....))..))))))))))))).))).))........ (-19.36 = -19.17 + -0.19)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:25:15 2006