
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 6,228,046 – 6,228,157 |
| Length | 111 |
| Max. P | 0.657698 |

| Location | 6,228,046 – 6,228,157 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 93.74 |
| Mean single sequence MFE | -24.28 |
| Consensus MFE | -20.28 |
| Energy contribution | -20.00 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.38 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.644647 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
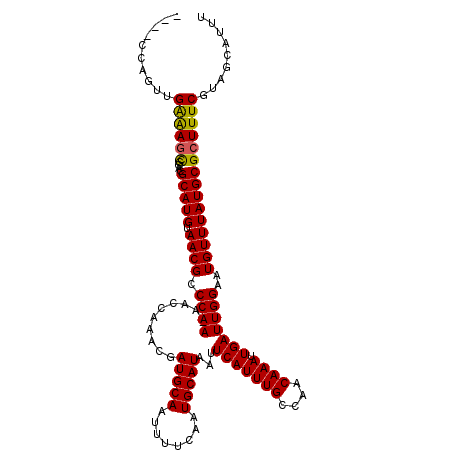
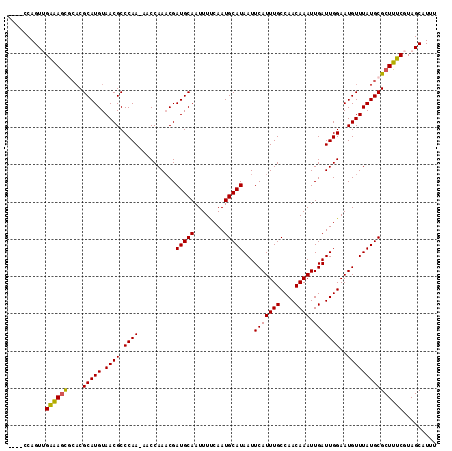
>2L_DroMel_CAF1 6228046 111 + 22407834 ----CCAGUUGAAAGCGCACGCAUGUAACGCCCAAAAACCAAACGAUGCAAUUUUCAAUGCAUAAUUCAUUUGCCAACAAAUUGAUUGGAAUGUUUAUGCGCUUUCGUAGCAUUU ----...((((((((((((.((.......)).......((((...(((((........)))))...(((((((....)))).)))))))........))))))))))..)).... ( -24.80) >DroSec_CAF1 64571 110 + 1 ----GCAGUUGAAAGCGCACGCAUGUAACGCCCAA-AACCAAACGAUGCAAUUUCCAAUGCAUAAUUCAUUUGCCAACAAAUUGAUUGGAAUGUUUAUGCGCUUUCGUAGCAUUU ----((...((((((((((.((((((.........-......)).))))...((((((((((.........)))(((....))))))))))......))))))))))..)).... ( -27.06) >DroSim_CAF1 77754 110 + 1 ----CCAGUUGAAAGCGCACGCAUGUAACGCCCAA-AACCAAACGAUGCAAUUUCCAAUGCAUAAUUCAUUUGCCAACAAAUUGAUUGGAAUGUUUAUGCGCUUUCGUAGCAUUU ----...((((((((((((.((((((.........-......)).))))...((((((((((.........)))(((....))))))))))......))))))))))..)).... ( -24.86) >DroEre_CAF1 61603 109 + 1 ----CCAGU-GAGAGUCCUCGCAUGUAACGCCCAA-AACCAAUCGAUGCAAUUUUCAAUGCAUAAUUCAUUUGCCAACAAAUUGAUUGGAAUGUUUAUGCGCUUUCGUAGCAUUU ----...((-(((....)))))((((.(((.....-..((((((((((((........))))).....(((((....))))))))))))...((....)).....))).)))).. ( -23.10) >DroYak_CAF1 76163 113 + 1 CCGCCCAGU-GGAAACCGACGCAUGUAACGCCCAA-AACCAAACGAUGCAAUUUUCAAUGCAUAAUUCAUUUGCCAACAAAUUGAUUGGAAUGUUUAUGCGCUUUCGUAGCAUUU ((((...))-))........(((((.((((.((((-.........(((((........)))))...(((((((....)))).)))))))..)))))))))(((.....))).... ( -21.60) >consensus ____CCAGUUGAAAGCGCACGCAUGUAACGCCCAA_AACCAAACGAUGCAAUUUUCAAUGCAUAAUUCAUUUGCCAACAAAUUGAUUGGAAUGUUUAUGCGCUUUCGUAGCAUUU ..........((((((....(((((.((((.((((..........(((((........)))))...(((((((....)))).)))))))..)))))))))))))))......... (-20.28 = -20.00 + -0.28)

| Location | 6,228,046 – 6,228,157 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 93.74 |
| Mean single sequence MFE | -29.84 |
| Consensus MFE | -23.14 |
| Energy contribution | -24.58 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.657698 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
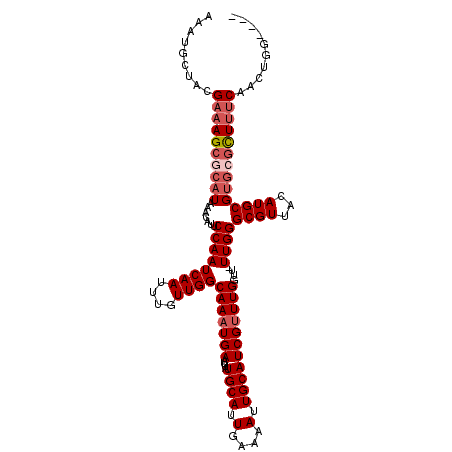
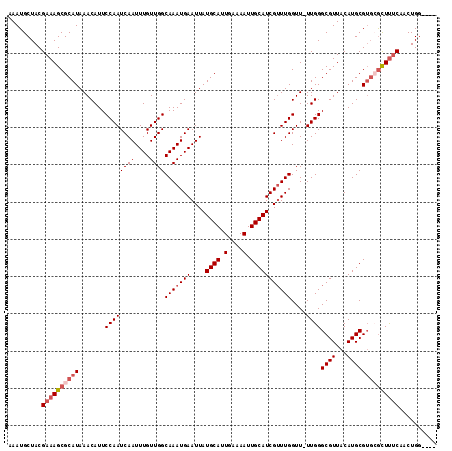
>2L_DroMel_CAF1 6228046 111 - 22407834 AAAUGCUACGAAAGCGCAUAAACAUUCCAAUCAAUUUGUUGGCAAAUGAAUUAUGCAUUGAAAAUUGCAUCGUUUGGUUUUUGGGCGUUACAUGCGUGCGCUUUCAACUGG---- .........((((((((((((((...(((((......)))))(((((((....((((.(....).)))))))))))))))....((((...))))))))))))))......---- ( -32.00) >DroSec_CAF1 64571 110 - 1 AAAUGCUACGAAAGCGCAUAAACAUUCCAAUCAAUUUGUUGGCAAAUGAAUUAUGCAUUGGAAAUUGCAUCGUUUGGUU-UUGGGCGUUACAUGCGUGCGCUUUCAACUGC---- .........((((((((((((((...(((((......)))))(((((((....((((.(....).))))))))))))))-)...((((...))))))))))))))......---- ( -32.00) >DroSim_CAF1 77754 110 - 1 AAAUGCUACGAAAGCGCAUAAACAUUCCAAUCAAUUUGUUGGCAAAUGAAUUAUGCAUUGGAAAUUGCAUCGUUUGGUU-UUGGGCGUUACAUGCGUGCGCUUUCAACUGG---- .........((((((((((((((...(((((......)))))(((((((....((((.(....).))))))))))))))-)...((((...))))))))))))))......---- ( -32.00) >DroEre_CAF1 61603 109 - 1 AAAUGCUACGAAAGCGCAUAAACAUUCCAAUCAAUUUGUUGGCAAAUGAAUUAUGCAUUGAAAAUUGCAUCGAUUGGUU-UUGGGCGUUACAUGCGAGGACUCUC-ACUGG---- ....((......(((((.....((..((((((((((..((....))..))))(((((.(....).))))).))))))..-.)).)))))....))(((....)))-.....---- ( -25.70) >DroYak_CAF1 76163 113 - 1 AAAUGCUACGAAAGCGCAUAAACAUUCCAAUCAAUUUGUUGGCAAAUGAAUUAUGCAUUGAAAAUUGCAUCGUUUGGUU-UUGGGCGUUACAUGCGUCGGUUUCC-ACUGGGCGG ..((((..(....).((((((.(((((((((......)))))..))))..))))))..........))))((((..((.-.(.(((((.....))))).).....-))..)))). ( -27.50) >consensus AAAUGCUACGAAAGCGCAUAAACAUUCCAAUCAAUUUGUUGGCAAAUGAAUUAUGCAUUGAAAAUUGCAUCGUUUGGUU_UUGGGCGUUACAUGCGUGCGCUUUCAACUGG____ .........((((((((((.......((((((((....))))(((((((....((((.(....).)))))))))))....))))((((...)))))))))))))).......... (-23.14 = -24.58 + 1.44)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:24:30 2006