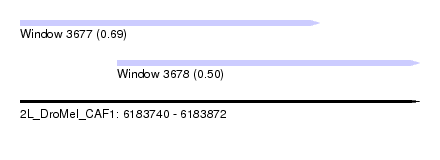

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 6,183,740 – 6,183,872 |
| Length | 132 |
| Max. P | 0.685128 |

| Location | 6,183,740 – 6,183,839 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 90.91 |
| Mean single sequence MFE | -30.76 |
| Consensus MFE | -27.66 |
| Energy contribution | -27.98 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.27 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.685128 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
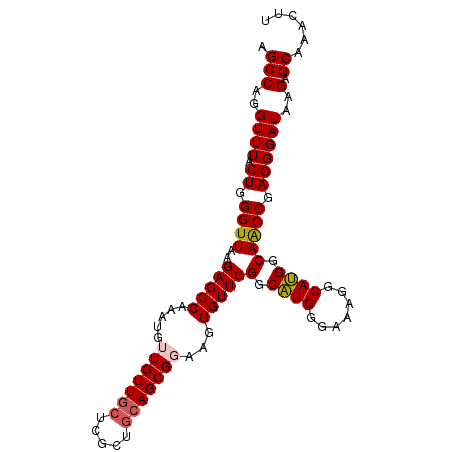
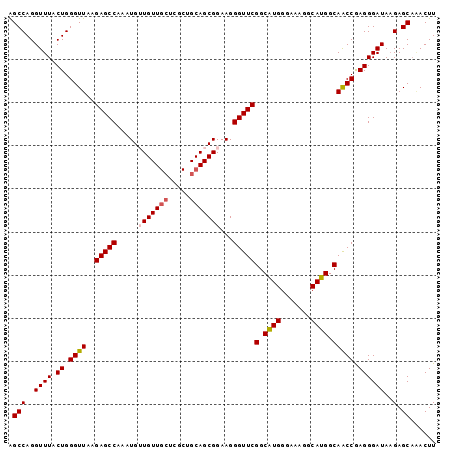
>2L_DroMel_CAF1 6183740 99 + 22407834 AGCCAGGUUUACUGGGUUAAGAGCCAAAUGUUGUUGCUCGCCGCAGCGGAAGGGUUCGGCAUGGGAAAGGCAUGGCAACCGAGGGAUAAGAGCAAACUU .(((..((((.((.((((..(((((.....(((((((.....)))))))...)))))(.((((.......)))).))))).))))))..).))...... ( -30.20) >DroSec_CAF1 20028 99 + 1 AGCCAGGUUUACUGGGUUAAGAGCCAAAUGUUGUUGCUCGCUGCAGCGGAAGGGUUCGGCAUGGGAAAGGCAUGGCAACCGAGGGAUAAGAGCAAACUU .(((..((((.((.((((..(((((.....(((((((.....)))))))...)))))(.((((.......)))).))))).))))))..).))...... ( -30.80) >DroSim_CAF1 29473 99 + 1 AGCCAGGUUUACUGGGUUAAGAGCCAAAUGUUGUUGCUCGCUGCAGCGGAAGGGUUCGGCAUGGGAAAGGCAUGGCAACCGAGGGAUAAGAGCAAACUU .(((..((((.((.((((..(((((.....(((((((.....)))))))...)))))(.((((.......)))).))))).))))))..).))...... ( -30.80) >DroEre_CAF1 19779 99 + 1 GGCCAGGUUUACUGGGUUAAGAGCCAAAUGGUGUUGCUCGCUGCAGCGGAAGGGUUCGGCCUGGGAAAAGCAGGGCAGCCAAGGGAUACGAGCCCACUU .(((..((((.(..(((...(((((...(..((((((.....))))))..).))))).)))..)...))))..)))......(((.......))).... ( -35.40) >DroYak_CAF1 30932 92 + 1 AGCCAGGUUUGCUGGGUUAAGAGCCAAAUGGUGUU-------GCAGCGGAAGGGUUCGGCAUGGGAAUGACAUGGCAACCAAGGGAUAAGAGCAGACUU ....(((((((((.(((.....)))......((((-------.(..(....)((((.(.((((.......)))).)))))..).))))..))))))))) ( -26.60) >consensus AGCCAGGUUUACUGGGUUAAGAGCCAAAUGUUGUUGCUCGCUGCAGCGGAAGGGUUCGGCAUGGGAAAGGCAUGGCAACCGAGGGAUAAGAGCAAACUU .(((..((((.((.((((..(((((.....(((((((.....)))))))...)))))(.((((.......)))).))))).))))))..).))...... (-27.66 = -27.98 + 0.32)

| Location | 6,183,772 – 6,183,872 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 88.50 |
| Mean single sequence MFE | -30.16 |
| Consensus MFE | -22.32 |
| Energy contribution | -23.00 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.74 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
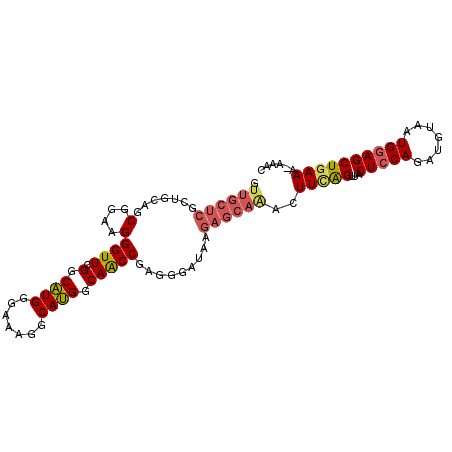
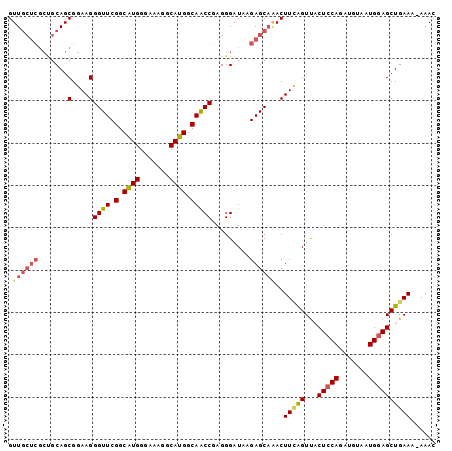
>2L_DroMel_CAF1 6183772 100 + 22407834 GUUGCUCGCCGCAGCGGAAGGGUUCGGCAUGGGAAAGGCAUGGCAACCGAGGGAUAAGAGCAAACUUAAGUCACUCCAGAUGUAAUGGAGCUGAAA-AAAC .((((((.((....(....)((((.(.((((.......)))).)))))...))....)))))).......((((((((.......))))).)))..-.... ( -29.60) >DroSec_CAF1 20060 101 + 1 GUUGCUCGCUGCAGCGGAAGGGUUCGGCAUGGGAAAGGCAUGGCAACCGAGGGAUAAGAGCAAACUUUGGUUACUCCAGAUGUAAUGGAGCUGAAUGAAAC (((((.....)))))......(((((((................((((((((............))))))))..((((.......)))))))))))..... ( -27.60) >DroSim_CAF1 29505 101 + 1 GUUGCUCGCUGCAGCGGAAGGGUUCGGCAUGGGAAAGGCAUGGCAACCGAGGGAUAAGAGCAAACUUUAGUCACUCCAGAUGUAAUGGAGCUGAAAGAAAC .((((((((....)).....((((.(.((((.......)))).))))).........))))))..(((..((((((((.......))))).)))..))).. ( -28.60) >DroEre_CAF1 19811 100 + 1 GUUGCUCGCUGCAGCGGAAGGGUUCGGCCUGGGAAAAGCAGGGCAGCCAAGGGAUACGAGCCCACUUCAGUUCCUGCAGAUGUAAUGGAGCUGAAA-AAAC (((((.....)))))....(((((((.(((.((....((...))..))..)))...)))))))..((((((((((((....)))..))))))))).-.... ( -37.70) >DroYak_CAF1 30964 94 + 1 GUU-------GCAGCGGAAGGGUUCGGCAUGGGAAUGACAUGGCAACCAAGGGAUAAGAGCAGACUUCAGUUCCUCCAGAUGUAAUGGAGCUGAAAAAAAC .((-------((..(....)((((.(.((((.......)))).)))))...........))))..(((((((((............)))))))))...... ( -27.30) >consensus GUUGCUCGCUGCAGCGGAAGGGUUCGGCAUGGGAAAGGCAUGGCAACCGAGGGAUAAGAGCAAACUUCAGUUACUCCAGAUGUAAUGGAGCUGAAA_AAAC .((((((.......(....)((((.(.((((.......)))).))))).........))))))..(((((...(((((.......))))))))))...... (-22.32 = -23.00 + 0.68)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:24:12 2006