
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 5,911,504 – 5,911,624 |
| Length | 120 |
| Max. P | 0.720206 |

| Location | 5,911,504 – 5,911,624 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.50 |
| Mean single sequence MFE | -30.98 |
| Consensus MFE | -21.19 |
| Energy contribution | -22.03 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.46 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.720206 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
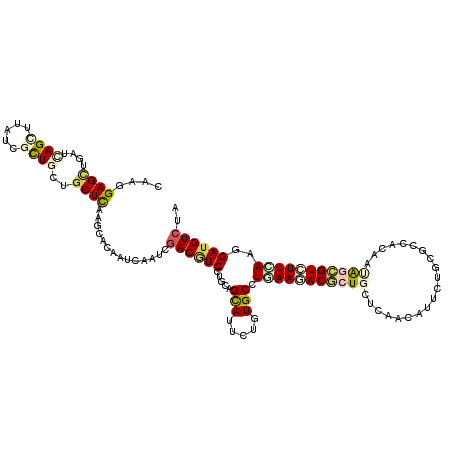
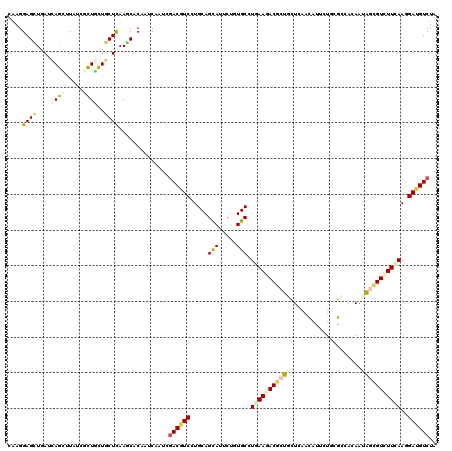
>2L_DroMel_CAF1 5911504 120 + 22407834 CAAGGAGCUGAUCAGCCUUUCGCUAUUGCUCAAGCACAACCAGUCGACGUCCUGCAGCAUUCUUUGCUUGAAGACACUGUUAAACAUUCUCCGGCACAACGCUGUAUUUAAAGAUGUCUA ....((((.(((.(((.....))))))))))..............((((((.((((((......((((.((.((...((.....)).)))).))))....))))))......)))))).. ( -24.60) >DroVir_CAF1 4029 120 + 1 CAAGGAGCUGAUCAGCUUAUCGCUGCUGCUCAAGCACAAUCAAUCGACGUCCUGCAGCAUCCUAUGCCUGAAGACGCUGCUCAACAUUCUGCGCCACAACAGCGUCUUCAAGGAUGUCUA ......(((((.((((........)))).)).)))..........(((((((((((........))).(((((((((((....................))))))))))))))))))).. ( -41.75) >DroPse_CAF1 1103 120 + 1 CAAGGAGUUGAUCAGCCUUUCGCUGCUCCUCAAGCACAAUCAGUCGACGUCCUGCAGUAUUCUGUGCCUCAAAACCCUCCUGAAUAUCCUACGGCACAAUGCAGUUUUCAAGGAUGUCUA ..(((((.....((((.....)))))))))...............((((((((((.((((..((((((........................)))))))))).)).....)))))))).. ( -29.66) >DroGri_CAF1 4094 120 + 1 UAAGGAGCUCAUCAGUUUGUCGCUACUGCUCAAGCACAAUCAAAGUACCUCAUGCAGCAUUCUGUGCCUCAAGACGCUGCUCAACAUAUUGCGCCACAAUAGCGUGUUCAAGGACGUCUA ...(((........(((((..((....)).))))).........((.(((...(((((..(((........))).)))))..........((((.......)))).....))))).))). ( -23.70) >DroWil_CAF1 1163 120 + 1 UAAGGAGUUGAUUAGUUUAUCUUUGCUGCUUAAACACAAUCAAUCGACAUCGUGCAGCAUUCUUUGCCUAAAGACACUCUUGAACAUUCUACGGCAUAAUAGUGUCUUUAAAGAUGUCUA ((((.(((.(((......)))...))).)))).............((((((.....(((.....))).((((((((((.(((..(........)..))).))))))))))..)))))).. ( -28.80) >DroMoj_CAF1 3941 120 + 1 CAAGGAGCUAAUCAGCUUAUCUCUGCUGCUCAAACACAAUCAGUCGACGUCGUGCAGCAUCCUCUGCAUGAAGACGCUGCUCAACAUUCUGCGCCACAAUAGCGUCUUCAAGGAUGUCUA ....((((.....(((........)))))))..............((((((((((((......))))))(((((((((((.((......)).))......)))))))))...)))))).. ( -37.40) >consensus CAAGGAGCUGAUCAGCUUAUCGCUGCUGCUCAAGCACAAUCAAUCGACGUCCUGCAGCAUUCUGUGCCUGAAGACGCUGCUCAACAUUCUGCGCCACAAUAGCGUCUUCAAGGAUGUCUA ....((((....((((.....))))..))))..............((((((.....(((.....))).(((((((((((....................)))))))))))..)))))).. (-21.19 = -22.03 + 0.84)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:21:47 2006