
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 5,686,122 – 5,686,217 |
| Length | 95 |
| Max. P | 0.972293 |

| Location | 5,686,122 – 5,686,217 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 87.25 |
| Mean single sequence MFE | -18.14 |
| Consensus MFE | -15.45 |
| Energy contribution | -16.34 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.69 |
| SVM RNA-class probability | 0.972293 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
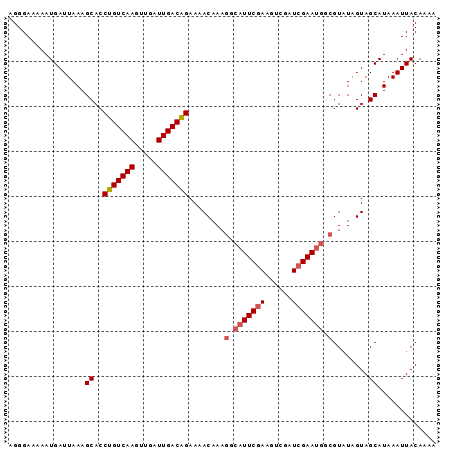
>2L_DroMel_CAF1 5686122 95 + 22407834 UGGGAAAAAUGAUUAAAGCACCUGUCAAGUUGAUUGACAGAAAACAAAGGCAUUCAAAGUCGAUCGAAUGGCGUAUAGUAGCAUAAAUUACAAAA .................((..(((((((.....)))))))........(.(((((..........))))).)........))............. ( -16.80) >DroSec_CAF1 131457 95 + 1 AGGGAAAAAUGAUUAAAGCACCUGUCAAGUUGAUUGACAGAAAACAAAGGCAUUCGAAGUCGAUCGAAUGGCGUAUGGUAGCAUAAAUUACAAAA .................((..(((((((.....)))))))....((..(.(((((((......))))))).)...))...))............. ( -21.60) >DroSim_CAF1 130910 95 + 1 AGGGAAAAAUGAUUAAAGCACCUGUCAAGUUGAUUGACAGAAAACAAAGGCAUUCGAAGUCGAUCGAAUGGCGUAUAGUAGCAUAAAUUACAAAA .................((..(((((((.....)))))))........(.(((((((......))))))).)........))............. ( -21.50) >DroEre_CAF1 134374 74 + 1 AGGGAAAAAUGAUUAAAGCACCUGUCAAGUUGAUUGACAGAA-ACAAA--------------------AGCCGUGUAGUAGCAUAAAUUACAAAA ........(((.(((..(((((((((((.....)))))))..-.....--------------------....))))..))))))........... ( -11.90) >DroYak_CAF1 132935 94 + 1 AGGGAAAAAUGAUUAAAGCACCUGUCAAGUUGAUUGACGGAA-ACAAAGCCAUUCGAAGUCGUUCGAAUGCCGUAUAGUAGCAUAAAUUACAAAA .................((..(((((((.....)))))))..-.....(.((((((((....)))))))).)........))............. ( -18.90) >consensus AGGGAAAAAUGAUUAAAGCACCUGUCAAGUUGAUUGACAGAAAACAAAGGCAUUCGAAGUCGAUCGAAUGGCGUAUAGUAGCAUAAAUUACAAAA .................((..(((((((.....)))))))........(.(((((((......))))))).)........))............. (-15.45 = -16.34 + 0.89)


| Location | 5,686,122 – 5,686,217 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 87.25 |
| Mean single sequence MFE | -16.16 |
| Consensus MFE | -12.63 |
| Energy contribution | -13.56 |
| Covariance contribution | 0.93 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.81 |
| SVM RNA-class probability | 0.857003 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
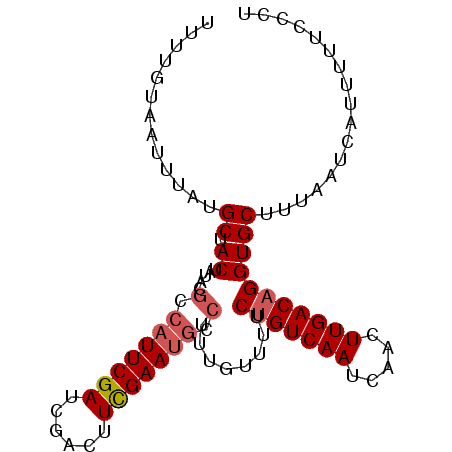
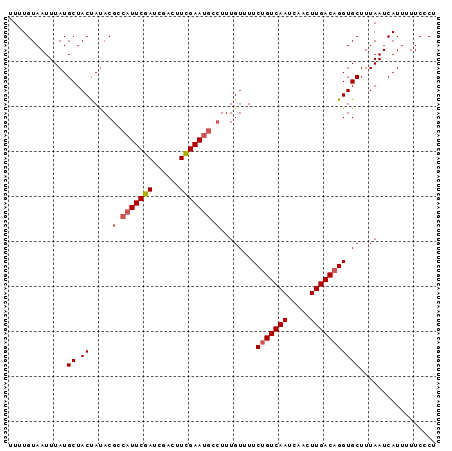
>2L_DroMel_CAF1 5686122 95 - 22407834 UUUUGUAAUUUAUGCUACUAUACGCCAUUCGAUCGACUUUGAAUGCCUUUGUUUUCUGUCAAUCAACUUGACAGGUGCUUUAAUCAUUUUUCCCA .............((.((.....(.(((((((......))))))).)........(((((((.....)))))))))))................. ( -15.60) >DroSec_CAF1 131457 95 - 1 UUUUGUAAUUUAUGCUACCAUACGCCAUUCGAUCGACUUCGAAUGCCUUUGUUUUCUGUCAAUCAACUUGACAGGUGCUUUAAUCAUUUUUCCCU .............((.(((....(.(((((((......))))))).).........((((((.....)))))))))))................. ( -18.60) >DroSim_CAF1 130910 95 - 1 UUUUGUAAUUUAUGCUACUAUACGCCAUUCGAUCGACUUCGAAUGCCUUUGUUUUCUGUCAAUCAACUUGACAGGUGCUUUAAUCAUUUUUCCCU .............((.((.....(.(((((((......))))))).)........(((((((.....)))))))))))................. ( -17.70) >DroEre_CAF1 134374 74 - 1 UUUUGUAAUUUAUGCUACUACACGGCU--------------------UUUGU-UUCUGUCAAUCAACUUGACAGGUGCUUUAAUCAUUUUUCCCU .............(((.......))).--------------------...((-.((((((((.....)))))))).))................. ( -11.10) >DroYak_CAF1 132935 94 - 1 UUUUGUAAUUUAUGCUACUAUACGGCAUUCGAACGACUUCGAAUGGCUUUGU-UUCCGUCAAUCAACUUGACAGGUGCUUUAAUCAUUUUUCCCU .............((.(((....(.((((((((....)))))))).).....-....(((((.....))))).)))))................. ( -17.80) >consensus UUUUGUAAUUUAUGCUACUAUACGCCAUUCGAUCGACUUCGAAUGCCUUUGUUUUCUGUCAAUCAACUUGACAGGUGCUUUAAUCAUUUUUCCCU .............((.((.....(.(((((((......))))))).)........(((((((.....)))))))))))................. (-12.63 = -13.56 + 0.93)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:19:47 2006