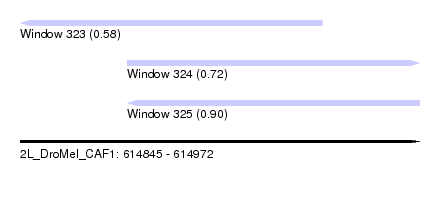
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 614,845 – 614,972 |
| Length | 127 |
| Max. P | 0.897115 |

| Location | 614,845 – 614,941 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 80.59 |
| Mean single sequence MFE | -25.20 |
| Consensus MFE | -13.94 |
| Energy contribution | -14.49 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.579140 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 614845 96 - 22407834 UUUCGGUGUGGUCCCCCCACUGAAACCCUCGCGAUCAAGCUCGAUCCCAUCAGGGGUUAAUACACUUGGCUGAUGUUUAAUGCUUAAUGGCAUUAA ((((((((.((....))))))))))......((..((((...((((((....))))))......))))..))....((((((((....)))))))) ( -27.60) >DroVir_CAF1 2600 78 - 1 UUUCGGUGUGGUCCCCCCCCCUACCC------------------CAGCGCCAGAGGUUAAUACACUUGCCUCAUGUUUAAUCCUAAAUGGCAUUAA ....((((.((.......)).)))).------------------....(((.(((((..........)))))..(((((....))))))))..... ( -17.00) >DroSim_CAF1 3351 96 - 1 UUUCGGUGUGGUCCCCCCACUGAAACCCUCGCGAUCAAGCUCGAUCCCAUCAGGGGUUAAUACACUUAGCUGAUGUUUAAUGCUUAAUGGCAUUAA ((((((((.((....)))))))))).......((((......)))).((((((((((......))))..)))))).((((((((....)))))))) ( -26.90) >DroEre_CAF1 6044 96 - 1 UUUCGGUGUGGUCCCCCCACUGAAACACUCGCGAUCAAGCUCGAUCCCAUCAGGGGUUAAUACACUUGGCUGAUGUUUAAUGCUUAAUGGCAUUAA ((((((((.((....)))))))))).((...((..((((...((((((....))))))......))))..))..))((((((((....)))))))) ( -27.90) >DroYak_CAF1 6579 96 - 1 UUUCGGUGUGGUCCCCCCACUGAAACCCUCGCGAUCAAGCUCGAUCCCAUCAGGGGUUAAUACACUUGGCUGAUGUUUAAUGCUUAAUGGCAUUAA ((((((((.((....))))))))))......((..((((...((((((....))))))......))))..))....((((((((....)))))))) ( -27.60) >DroMoj_CAF1 2349 78 - 1 UUUCGGUGUGGUCCCUCGCCUCACCG------------------CCCCGCCAGGGGUUAAUACACUUGCCUCAUGUUUAAUCCUAAAUGGCAUUAA ...(((((.(((.....))).)))))------------------....((((((((((((.(((.........)))))))))))...))))..... ( -24.20) >consensus UUUCGGUGUGGUCCCCCCACUGAAACCCUCGCGAUCAAGCUCGAUCCCAUCAGGGGUUAAUACACUUGGCUGAUGUUUAAUGCUUAAUGGCAUUAA ....((((((...((((.............((......))............))))....))))))..........((((((((....)))))))) (-13.94 = -14.49 + 0.56)

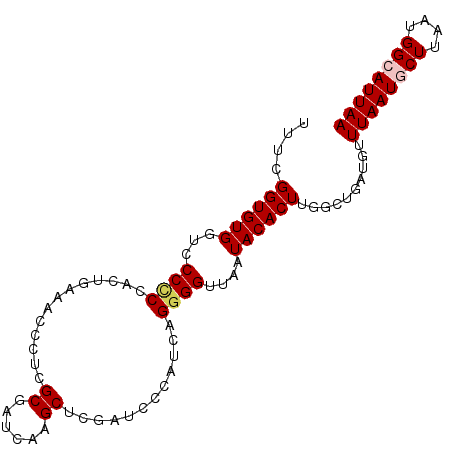
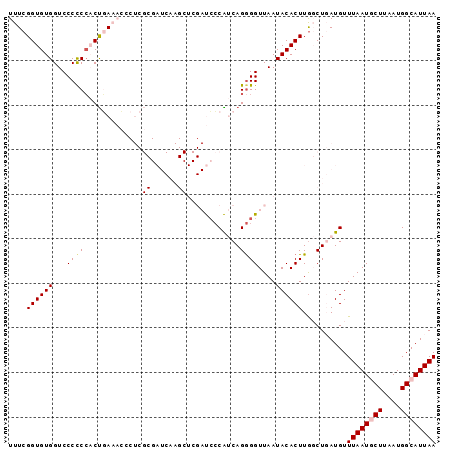
| Location | 614,879 – 614,972 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 99.57 |
| Mean single sequence MFE | -34.02 |
| Consensus MFE | -33.52 |
| Energy contribution | -33.72 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.99 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.716039 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
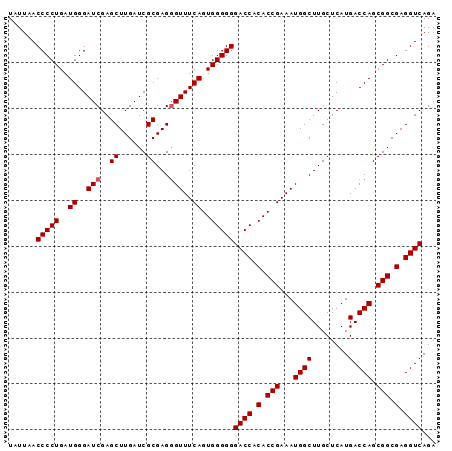
>2L_DroMel_CAF1 614879 93 + 22407834 UAUUAACCCCUGAUGGGAUCGAGCUUGAUCGCGAGGGUUUCAGUGGGGGGACCACACCGAAAUGGCUUGCUCAUGACCAGCGGCGAGGUCAGA ......(((((..((..(((..((......))...)))..))..)))))((((.(.(((...((((........).))).))).).))))... ( -34.20) >DroSec_CAF1 4247 93 + 1 UAUUAACCCCUGAUGGGAUCGAGCUUGAUCGCGAGGGUUUCAGUGGGGGGACCACACCGAAAUGGCUUGCUCAUGACCAGCGGCGAGGUCAGA ......(((((..((..(((..((......))...)))..))..)))))((((.(.(((...((((........).))).))).).))))... ( -34.20) >DroSim_CAF1 3385 93 + 1 UAUUAACCCCUGAUGGGAUCGAGCUUGAUCGCGAGGGUUUCAGUGGGGGGACCACACCGAAAUGGCUUGCUCAUGACCAGCGGCGAGGUCAGA ......(((((..((..(((..((......))...)))..))..)))))((((.(.(((...((((........).))).))).).))))... ( -34.20) >DroEre_CAF1 6078 93 + 1 UAUUAACCCCUGAUGGGAUCGAGCUUGAUCGCGAGUGUUUCAGUGGGGGGACCACACCGAAAUGGCUUGCUCAUGACCAGCGGCGAGGUCAGA .....(((((((.((((((((....)))))(((((((((((.(((((....)).))).))))).))))))......))).))).).))).... ( -33.30) >DroYak_CAF1 6613 93 + 1 UAUUAACCCCUGAUGGGAUCGAGCUUGAUCGCGAGGGUUUCAGUGGGGGGACCACACCGAAAUGGCUUGCUCAUGACCAGCGGCGAGGUCAGA ......(((((..((..(((..((......))...)))..))..)))))((((.(.(((...((((........).))).))).).))))... ( -34.20) >consensus UAUUAACCCCUGAUGGGAUCGAGCUUGAUCGCGAGGGUUUCAGUGGGGGGACCACACCGAAAUGGCUUGCUCAUGACCAGCGGCGAGGUCAGA ......(((((..((..(((..((......))...)))..))..)))))((((.(.(((...((((........).))).))).).))))... (-33.52 = -33.72 + 0.20)

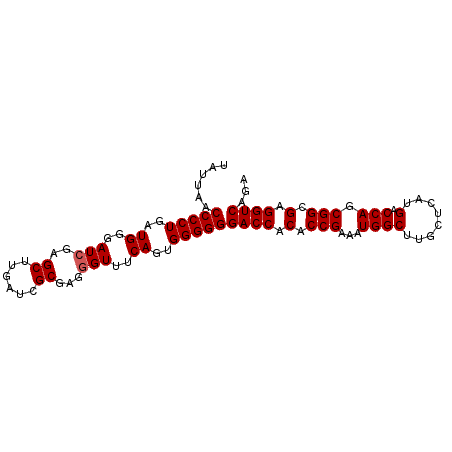
| Location | 614,879 – 614,972 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 99.57 |
| Mean single sequence MFE | -29.40 |
| Consensus MFE | -29.40 |
| Energy contribution | -29.40 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.50 |
| Structure conservation index | 1.00 |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.897115 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
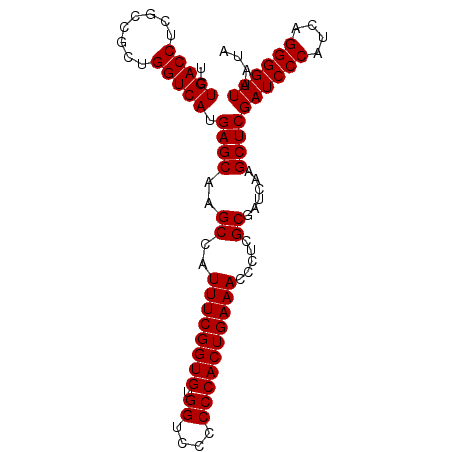
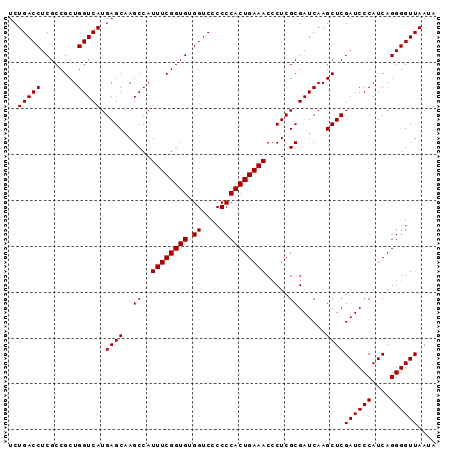
>2L_DroMel_CAF1 614879 93 - 22407834 UCUGACCUCGCCGCUGGUCAUGAGCAAGCCAUUUCGGUGUGGUCCCCCCACUGAAACCCUCGCGAUCAAGCUCGAUCCCAUCAGGGGUUAAUA ..(((((........))))).((((..((..((((((((.((....)))))))))).....))......))))((((((....)))))).... ( -29.40) >DroSec_CAF1 4247 93 - 1 UCUGACCUCGCCGCUGGUCAUGAGCAAGCCAUUUCGGUGUGGUCCCCCCACUGAAACCCUCGCGAUCAAGCUCGAUCCCAUCAGGGGUUAAUA ..(((((........))))).((((..((..((((((((.((....)))))))))).....))......))))((((((....)))))).... ( -29.40) >DroSim_CAF1 3385 93 - 1 UCUGACCUCGCCGCUGGUCAUGAGCAAGCCAUUUCGGUGUGGUCCCCCCACUGAAACCCUCGCGAUCAAGCUCGAUCCCAUCAGGGGUUAAUA ..(((((........))))).((((..((..((((((((.((....)))))))))).....))......))))((((((....)))))).... ( -29.40) >DroEre_CAF1 6078 93 - 1 UCUGACCUCGCCGCUGGUCAUGAGCAAGCCAUUUCGGUGUGGUCCCCCCACUGAAACACUCGCGAUCAAGCUCGAUCCCAUCAGGGGUUAAUA ..(((((........))))).((((..((..((((((((.((....)))))))))).....))......))))((((((....)))))).... ( -29.40) >DroYak_CAF1 6613 93 - 1 UCUGACCUCGCCGCUGGUCAUGAGCAAGCCAUUUCGGUGUGGUCCCCCCACUGAAACCCUCGCGAUCAAGCUCGAUCCCAUCAGGGGUUAAUA ..(((((........))))).((((..((..((((((((.((....)))))))))).....))......))))((((((....)))))).... ( -29.40) >consensus UCUGACCUCGCCGCUGGUCAUGAGCAAGCCAUUUCGGUGUGGUCCCCCCACUGAAACCCUCGCGAUCAAGCUCGAUCCCAUCAGGGGUUAAUA ..(((((........))))).((((..((..((((((((.((....)))))))))).....))......))))((((((....)))))).... (-29.40 = -29.40 + 0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:29:42 2006