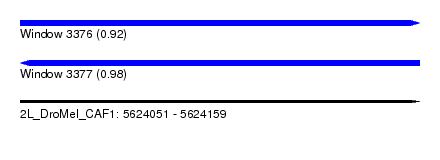

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 5,624,051 – 5,624,159 |
| Length | 108 |
| Max. P | 0.979808 |

| Location | 5,624,051 – 5,624,159 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 80.99 |
| Mean single sequence MFE | -25.00 |
| Consensus MFE | -19.45 |
| Energy contribution | -20.23 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.78 |
| SVM decision value | 1.13 |
| SVM RNA-class probability | 0.919348 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
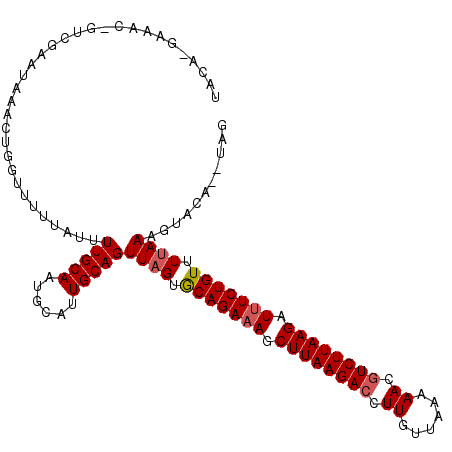
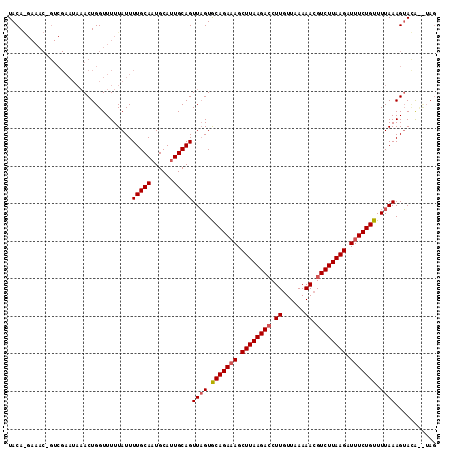
>2L_DroMel_CAF1 5624051 108 + 22407834 UACAGGAAACGGUCGAAUAAACUGGUUUUUAUUUUGCAAUGCAAUGCAGUUUGUGCAGACAGCUUAAGACCUUGUUAAAAA-GUCUUAAGAUUUCUGUUUUAAAGAACA----- .(((((((...(((..((((((((.........((((...))))..))))))))...)))..((((((((.((.....)).-)))))))).)))))))...........----- ( -25.90) >DroSec_CAF1 68046 114 + 1 UACAAGAAACUGUCGAAUAAACUGGUUUUUAUUUUGCAAUGCAUUGCAGUUAGUACAGAAAGCUUAAGACCUUGUUAAAAACGUCUUAAGAUUUCUGUUUUAAAGUAUGCCUAG ..........(((.(((((((......))))))).)))..((((.((..((((.(((((((.((((((((.((......)).)))))))).))))))).)))).)))))).... ( -27.50) >DroSim_CAF1 69809 98 + 1 UACA----------------ACUGGUUUUUAUUUUGCAAUGCAUUGCAGUUAGUGCAGAAAGCUUAAGAGCUUGUUAAAAACGUCUUAAGAUUUCUGUUUUAAAGUAGAUUUAG ....----------------.((((..((((((((((((....))))))((((.(((((((.(((((((..((......))..))))))).))))))).)))))))))).)))) ( -21.60) >consensus UACA_GAAAC_GUCGAAUAAACUGGUUUUUAUUUUGCAAUGCAUUGCAGUUAGUGCAGAAAGCUUAAGACCUUGUUAAAAACGUCUUAAGAUUUCUGUUUUAAAGUACA__UAG .................................(((((......)))))((((.(((((((.((((((((.((......)).)))))))).))))))).))))........... (-19.45 = -20.23 + 0.78)

| Location | 5,624,051 – 5,624,159 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 80.99 |
| Mean single sequence MFE | -21.17 |
| Consensus MFE | -16.33 |
| Energy contribution | -17.00 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.49 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.85 |
| SVM RNA-class probability | 0.979808 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 5624051 108 - 22407834 -----UGUUCUUUAAAACAGAAAUCUUAAGAC-UUUUUAACAAGGUCUUAAGCUGUCUGCACAAACUGCAUUGCAUUGCAAAAUAAAAACCAGUUUAUUCGACCGUUUCCUGUA -----...........((((((((((((((((-(((.....)))))))))))..(((((((.....))))((((...))))...................))).)))).)))). ( -20.50) >DroSec_CAF1 68046 114 - 1 CUAGGCAUACUUUAAAACAGAAAUCUUAAGACGUUUUUAACAAGGUCUUAAGCUUUCUGUACUAACUGCAAUGCAUUGCAAAAUAAAAACCAGUUUAUUCGACAGUUUCUUGUA .(((((..........(((((((.((((((((.((......)).)))))))).)))))))......(((((....)))))............)))))....((((....)))). ( -24.70) >DroSim_CAF1 69809 98 - 1 CUAAAUCUACUUUAAAACAGAAAUCUUAAGACGUUUUUAACAAGCUCUUAAGCUUUCUGCACUAACUGCAAUGCAUUGCAAAAUAAAAACCAGU----------------UGUA .................((((((.(((((((.((((.....))))))))))).)))))).......(((((....)))))..............----------------.... ( -18.30) >consensus CUA__UAUACUUUAAAACAGAAAUCUUAAGACGUUUUUAACAAGGUCUUAAGCUUUCUGCACUAACUGCAAUGCAUUGCAAAAUAAAAACCAGUUUAUUCGAC_GUUUC_UGUA .................((((((.((((((((.((......)).)))))))).)))))).......((((......)))).................................. (-16.33 = -17.00 + 0.67)
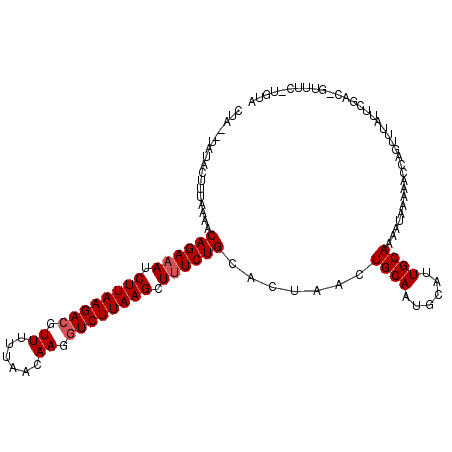



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:19:19 2006