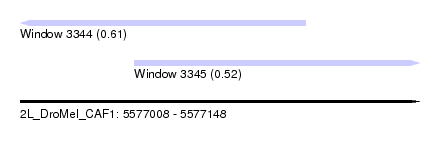
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 5,577,008 – 5,577,148 |
| Length | 140 |
| Max. P | 0.611108 |

| Location | 5,577,008 – 5,577,108 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.59 |
| Mean single sequence MFE | -20.77 |
| Consensus MFE | -14.50 |
| Energy contribution | -14.18 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.611108 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
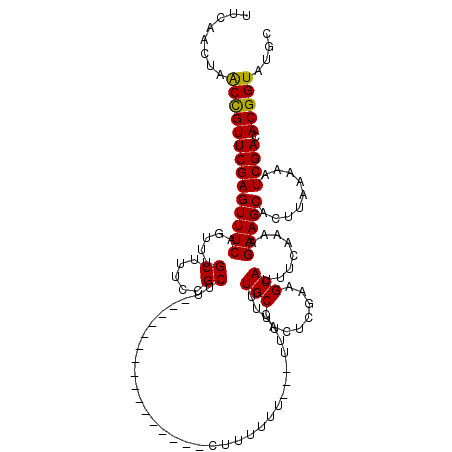
>2L_DroMel_CAF1 5577008 100 - 22407834 UUCAACUAACCGUUCGAGUUUCAGUUUGCUUUCCGCUU----------------UUUUUU----UUUUGUUGCCACUCGAAGCAUUUCAAAAGAAGCACUUAAAAAUCGACACGGUAUGC ........(((((((((((........)).....((((----------------(((((.----...((((.........))))....))))))))).........)))).))))).... ( -20.20) >DroSec_CAF1 21039 104 - 1 AUCAACUAACCGUUCGAGUUUCAGUUUGCUUUCCGCUU----------------CUUUUUUUUUUUUUUUUGCCACUCGAAGCAUUUCAAAAGAAGCACUUAAAAAUCGACACGGUAUGC ........(((((((((((........)).....((((----------------(((((...........(((........)))....))))))))).........)))).))))).... ( -19.86) >DroSim_CAF1 22690 101 - 1 UUCAACUAACCGUUCGAGUUUCAGUUUGCUUUCCGCUU----------------CUUUUUU---UUGUUUUGCCACUCGAAGCAUUUCAAAAGAAGCUCUUAAAAAUCGACACGGUAUGC ........(((((((((((........)).....((((----------------(((((..---.(((((((.....)))))))....))))))))).........)))).))))).... ( -25.40) >DroEre_CAF1 20961 117 - 1 UCCAACUUGCCGUUCGAGUUUCAGUUUGCUUUCCGCCUGGUUUUUACUUUUUUACUUUUUU---UUUUUUUGCCACACCGAGCAUUUCAAAAGAAGCACUUAAAAAUCGACACGGUAUGC .......((((((((((...((((...((.....))))))........(((((((((((((---(.....(((........)))....))))))))....)))))))))).))))))... ( -20.00) >DroYak_CAF1 20917 91 - 1 -CCAACUAACUGUUCGAGUUUCAGUUUGCUUUCCGC----------------------------UUUUUUUGCCACUGCUAGCAUUUCAAAAGAAGCACUUAAAAAUCGACACGGUAUGC -.......(((((((((((........)).....((----------------------------((((((((....((....))...)))))))))).........)))).))))).... ( -18.40) >consensus UUCAACUAACCGUUCGAGUUUCAGUUUGCUUUCCGCUU________________CUUUUUU___UUUUUUUGCCACUCGAAGCAUUUCAAAAGAAGCACUUAAAAAUCGACACGGUAUGC ........((((((((((((((.....((.....))..................................(((........)))........))))).........)))).))))).... (-14.50 = -14.18 + -0.32)

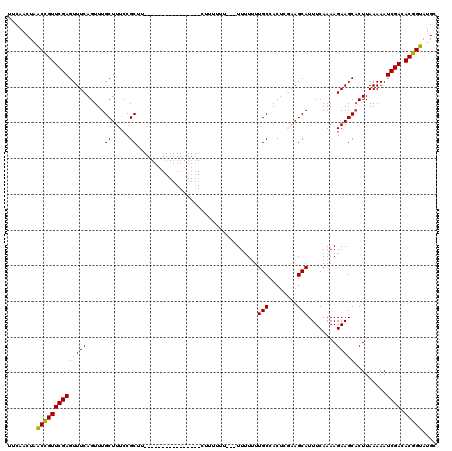
| Location | 5,577,048 – 5,577,148 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 84.53 |
| Mean single sequence MFE | -27.12 |
| Consensus MFE | -20.88 |
| Energy contribution | -22.25 |
| Covariance contribution | 1.38 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.77 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.516965 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
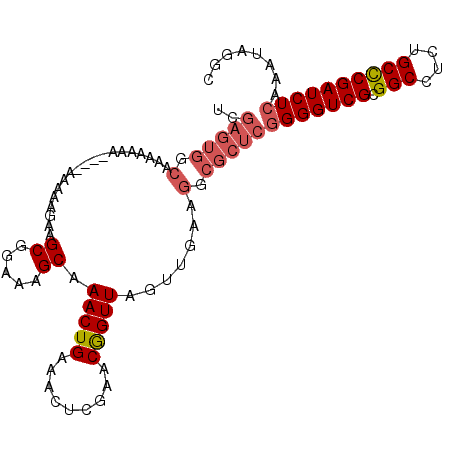
>2L_DroMel_CAF1 5577048 100 + 22407834 UCGAGUGGCAACAAAA----AAAAAAAAGCGGAAAGCAAACUGAAACUCGAACGGUUAGUUGAAGGCGCUCGGGGUCGCGGCCUCUGCCCGAUCUCAAAUAGGC ..(((((.(..(((..----........((.....)).(((((.........)))))..)))..).)))))(((((((.(((....))))))))))........ ( -29.30) >DroSec_CAF1 21079 104 + 1 UCGAGUGGCAAAAAAAAAAAAAAAAGAAGCGGAAAGCAAACUGAAACUCGAACGGUUAGUUGAUGGCGCUCGGGGUCGCGGCCUCUGCCCGAUCUCAAAUAGGC ..(((((.((..................((.....)).(((((.........)))))......)).)))))(((((((.(((....))))))))))........ ( -29.00) >DroSim_CAF1 22730 101 + 1 UCGAGUGGCAAAACAA---AAAAAAGAAGCGGAAAGCAAACUGAAACUCGAACGGUUAGUUGAAGGCGCUCGGGGUCGCGGCCUCUGCCCGAUCUCAAAUAGGC ..(((((.(..(((..---.........((.....)).(((((.........))))).)))...).)))))(((((((.(((....))))))))))........ ( -29.40) >DroYak_CAF1 20957 83 + 1 AGCAGUGGCAAAAAAA------------GCGGAAAGCAAACUGAAACUCGAACAGUUAGUUGG---------GGGUCGCGGCCCCUGCUCGAUCUCAAAUAGGC (((((.(((.......------------((((..(((.(((((.........))))).)))..---------...)))).))).)))))............... ( -20.80) >consensus UCGAGUGGCAAAAAAA____AAAAAGAAGCGGAAAGCAAACUGAAACUCGAACGGUUAGUUGAAGGCGCUCGGGGUCGCGGCCUCUGCCCGAUCUCAAAUAGGC ..(((((.(...................((.....)).(((((.........))))).......).)))))(((((((.(((....))))))))))........ (-20.88 = -22.25 + 1.38)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:18:47 2006