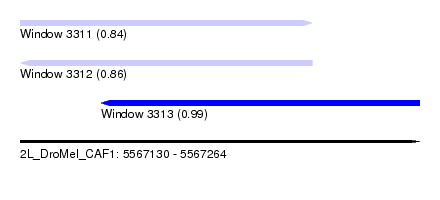
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 5,567,130 – 5,567,264 |
| Length | 134 |
| Max. P | 0.992004 |

| Location | 5,567,130 – 5,567,228 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 95.31 |
| Mean single sequence MFE | -18.87 |
| Consensus MFE | -17.52 |
| Energy contribution | -17.12 |
| Covariance contribution | -0.40 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.29 |
| Structure conservation index | 0.93 |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.843382 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 5567130 98 + 22407834 CUUCACCUUUCACCUCUUUUGUUUUUCUUUUCCGCUCUUUUUCCAUUGAGCCGAAGACACACACUCAAAAAGCGUGAGUCGCACAAUGGGUCUUGUUG ...........((((.((.(((.......(((.((((..........)))).)))(((.(((.((.....)).))).)))))).)).))))....... ( -19.20) >DroSec_CAF1 11807 98 + 1 CUUCACCUUUCAUCUCUUUUGUUUUUCUUUUCCGCUCUUUUUCCAUUGAGCCGAAGACACACACUCAAAAAGCGUGAGUCGCACAAUGGGACUCGUUG ............((((..((((.......(((.((((..........)))).)))(((.(((.((.....)).))).)))..)))).))))....... ( -19.10) >DroSim_CAF1 11754 98 + 1 CUUCACCUUUCAUCUCUUUUGUUUUUCUUUUCCGCUCUUUUUCCAUUGAGCCGAAGACACACACUCAAAAAGCGUGAGUCGCACAAUGGGACUCGUUG ............((((..((((.......(((.((((..........)))).)))(((.(((.((.....)).))).)))..)))).))))....... ( -19.10) >DroEre_CAF1 11798 97 + 1 CUUCACCUUUCACCUCCUUUGUUUUUCUUUUCCGCUCUUUUUCCAUCGAGCUGAAGACACACACUCAAAAAGCGUGAGUCGCACAAUGGGACUCG-UG ..........(((.((((((((.......(((.((((..........)))).)))(((.(((.((.....)).))).)))..)))).))))...)-)) ( -19.80) >DroYak_CAF1 12003 98 + 1 CUUCACCUUUCACCUCCUUUGUUUUUCUUUUCCGCUCUUUUUCCAUUGAGUUGAAGACACACACUCAAAAAGCGUGAGUCCCACAAUGGGUCUUGUUG .........((((((..((((......(((((.((((..........)))).)))))........)))).)).))))(.(((.....))).)...... ( -17.14) >consensus CUUCACCUUUCACCUCUUUUGUUUUUCUUUUCCGCUCUUUUUCCAUUGAGCCGAAGACACACACUCAAAAAGCGUGAGUCGCACAAUGGGACUCGUUG ..............(((.((((.......(((.((((..........)))).)))(((.(((.((.....)).))).)))..)))).)))........ (-17.52 = -17.12 + -0.40)



| Location | 5,567,130 – 5,567,228 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 95.31 |
| Mean single sequence MFE | -24.01 |
| Consensus MFE | -20.42 |
| Energy contribution | -20.46 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.81 |
| SVM RNA-class probability | 0.855457 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 5567130 98 - 22407834 CAACAAGACCCAUUGUGCGACUCACGCUUUUUGAGUGUGUGUCUUCGGCUCAAUGGAAAAAGAGCGGAAAAGAAAAACAAAAGAGGUGAAAGGUGAAG .......((((.((((..(((.(((((((...))))))).)))(((.((((..(....)..)))).))).......))))..).)))........... ( -23.60) >DroSec_CAF1 11807 98 - 1 CAACGAGUCCCAUUGUGCGACUCACGCUUUUUGAGUGUGUGUCUUCGGCUCAAUGGAAAAAGAGCGGAAAAGAAAAACAAAAGAGAUGAAAGGUGAAG ....(((((.(.....).))))).((((((((.....(((.(((((.((((..(....)..)))).))..)))...)))........))))))))... ( -23.62) >DroSim_CAF1 11754 98 - 1 CAACGAGUCCCAUUGUGCGACUCACGCUUUUUGAGUGUGUGUCUUCGGCUCAAUGGAAAAAGAGCGGAAAAGAAAAACAAAAGAGAUGAAAGGUGAAG ....(((((.(.....).))))).((((((((.....(((.(((((.((((..(....)..)))).))..)))...)))........))))))))... ( -23.62) >DroEre_CAF1 11798 97 - 1 CA-CGAGUCCCAUUGUGCGACUCACGCUUUUUGAGUGUGUGUCUUCAGCUCGAUGGAAAAAGAGCGGAAAAGAAAAACAAAGGAGGUGAAAGGUGAAG ((-((((((.(.....).))))).((((((((...(((...((((..((((..(....)..))))....))))...))).))))))))....)))... ( -28.10) >DroYak_CAF1 12003 98 - 1 CAACAAGACCCAUUGUGGGACUCACGCUUUUUGAGUGUGUGUCUUCAACUCAAUGGAAAAAGAGCGGAAAAGAAAAACAAAGGAGGUGAAAGGUGAAG ...((...(((.((((.((((.(((((((...))))))).))))((..(((..(....)..)))..))........)))).)).).)).......... ( -21.10) >consensus CAACGAGUCCCAUUGUGCGACUCACGCUUUUUGAGUGUGUGUCUUCGGCUCAAUGGAAAAAGAGCGGAAAAGAAAAACAAAAGAGGUGAAAGGUGAAG .........((((((.(((((.(((((((...))))))).)))....)).)))))).......................................... (-20.42 = -20.46 + 0.04)

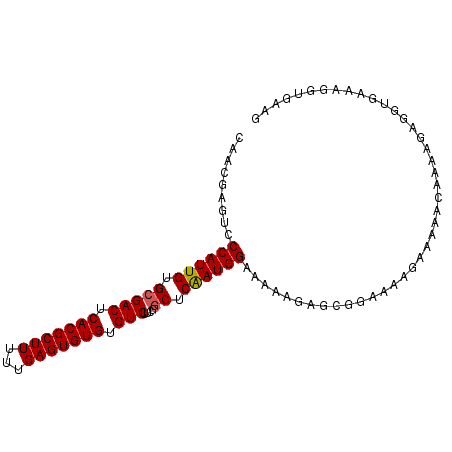
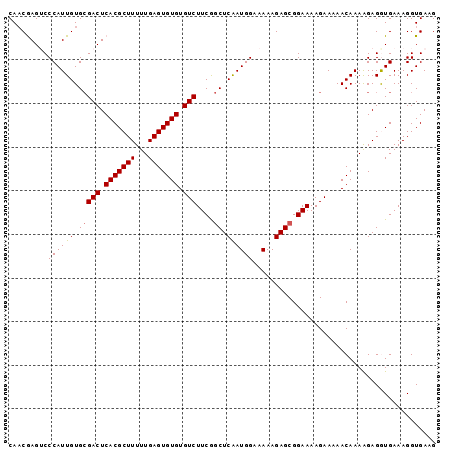
| Location | 5,567,157 – 5,567,264 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 88.07 |
| Mean single sequence MFE | -27.22 |
| Consensus MFE | -24.42 |
| Energy contribution | -24.54 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.90 |
| SVM decision value | 2.30 |
| SVM RNA-class probability | 0.992004 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 5567157 107 - 22407834 UCUA--AUAAUUUCAUAAUAUUUCUACAC--AAUAUUUGGCAACAAGACCCAUUGUGCGACUCACGCUUUUUGAGUGUGUGUCUUCGGCUCAAUGGAAAAAGAGCGGAAAA ....--..............((((..(..--....((((....))))..((((((.(((((.(((((((...))))))).)))....)).)))))).......)..)))). ( -28.50) >DroSec_CAF1 11834 109 - 1 UUUAAAAACACUUUACAAUAUUUCAACAC--AAUAUUUCGCAACGAGUCCCAUUGUGCGACUCACGCUUUUUGAGUGUGUGUCUUCGGCUCAAUGGAAAAAGAGCGGAAAA .............................--....((((((........((((((.(((((.(((((((...))))))).)))....)).)))))).......)))))).. ( -26.56) >DroSim_CAF1 11781 111 - 1 UUUACAAAAACUUCAUAAUAUUUUAAAACUAAAUAUUUCGCAACGAGUCCCAUUGUGCGACUCACGCUUUUUGAGUGUGUGUCUUCGGCUCAAUGGAAAAAGAGCGGAAAA ...................................((((((........((((((.(((((.(((((((...))))))).)))....)).)))))).......)))))).. ( -26.56) >DroEre_CAF1 11825 108 - 1 UUUACAGUAAUUUUACAAUAUGUGAACGC--AAUAUUUCGCA-CGAGUCCCAUUGUGCGACUCACGCUUUUUGAGUGUGUGUCUUCAGCUCGAUGGAAAAAGAGCGGAAAA ......((...(((((.....))))).))--....((((((.-(.....((((((.(((((.(((((((...))))))).)))....)).)))))).....).)))))).. ( -29.80) >DroYak_CAF1 12030 109 - 1 UGUACAAAAAUUUUGCAAUAUUUUAACAC--AACAUUUUGCAACAAGACCCAUUGUGGGACUCACGCUUUUUGAGUGUGUGUCUUCAACUCAAUGGAAAAAGAGCGGAAAA ..........((((((.............--....(((((...))))).((((((.(((((.(((((((...))))))).))))).....)))))).......)))))).. ( -24.70) >consensus UUUACAAAAAUUUUACAAUAUUUCAACAC__AAUAUUUCGCAACGAGUCCCAUUGUGCGACUCACGCUUUUUGAGUGUGUGUCUUCGGCUCAAUGGAAAAAGAGCGGAAAA ...................................((((((........((((((.(((((.(((((((...))))))).)))....)).)))))).......)))))).. (-24.42 = -24.54 + 0.12)
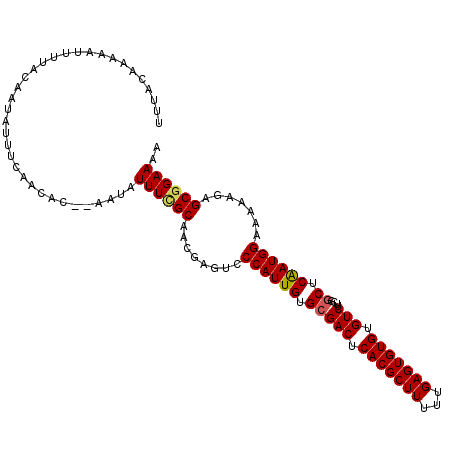



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:18:16 2006