
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 5,533,642 – 5,533,760 |
| Length | 118 |
| Max. P | 0.832082 |

| Location | 5,533,642 – 5,533,760 |
|---|---|
| Length | 118 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 77.49 |
| Mean single sequence MFE | -32.68 |
| Consensus MFE | -19.17 |
| Energy contribution | -17.87 |
| Covariance contribution | -1.30 |
| Combinations/Pair | 1.41 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.59 |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.746144 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
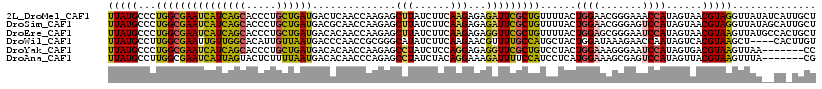
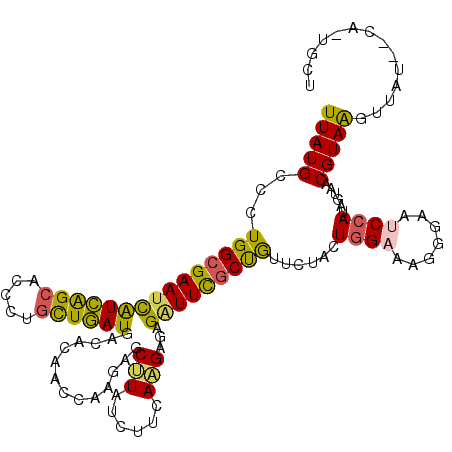
>2L_DroMel_CAF1 5533642 118 + 22407834 UUAUGCCCUGGCGAAUCAUCAGCACCCUGCUGAUGACUCAACCAAGAGCUUAUCUUCAAGAGAGAUUCGCUGUUUUACUGGAACGGGAAACCAUAGUAACGUAGGUUAUAUCAUUGCU ....((....))((.((((((((.....)))))))).))((((((.(((..(((((.....)))))..))).))((((((....((....)).))))))....))))........... ( -36.00) >DroSim_CAF1 7670 118 + 1 UUAUGCCCUGGCGAAUCAUCAGCACCCUGCUGAUGACGCAACCAAGAGCUUAUCUUCAAGAGAGAUUCGCUGUUUUACUGGAACGGGAGUCCAUAGUAACGUAGGUUAUAGCAUUGCU ..((((....(((..((((((((.....)))))))))))((((((.(((..(((((.....)))))..))).))((((((((.(....))))..)))))....))))...)))).... ( -35.60) >DroEre_CAF1 7722 118 + 1 UUAUGCCCUGGCGAAUCAUCAGCACCCUGCUGAUGACACAACCAAGAGCUUAUCUUCAAGAGAGGUUCGCUGUUUUACUGGAGCGGGAAUCCAUAGUAACGUAAGUUAUGCCACUGCU ....((..((((...((((((((.....)))))))).....((..((((((.((.....)).))))))(((.(......).))))).........(((((....)))))))))..)). ( -38.00) >DroWil_CAF1 7795 114 + 1 UUAUGCCUUGGCGAAUUGUUGGCACAUUGUUAAUGACCCAACCGCGGGCAUAUCUUCAAGAACGUUUUGCCAUGCUACUGGAUAAAGAACCAAUAGUCACGUAAGCU----CACUUGU .(((((((..(((....(((((..(((.....)))..)))))))))))))))....((((...((((.((...((((.(((........))).))))...)))))).----..)))). ( -27.00) >DroYak_CAF1 7740 111 + 1 UUAUGCCCUGGCGAAUCAUCAGCACCCUGCUGAUGACACAACCAAGAGCCUAUCUCCAGGAGAGGUUCGCUGUCCUACUGGAAAGGGAAUCCAUAGUGACGUAAGUUAA-------CC ((((((((((((...((((((((.....)))))))).........((((((.((.....)).))))))))).(((....))).))))....)))))((((....)))).-------.. ( -37.50) >DroAna_CAF1 7420 111 + 1 UUAUGCCUUGGCGAAUCAUUAGUACUCUUUUAAUGACACAACCCAGAGCCUAUCUACAGGAAAGAUUUUCCAUCCUCAUGGAAAGCGAGUCCAUAGUUACGUAAGUUUA-------CG ....((.((((....(((((((.......)))))))......)))).)).........(((..(.((((((((....)))))))))...)))...((.((....))..)-------). ( -22.00) >consensus UUAUGCCCUGGCGAAUCAUCAGCACCCUGCUGAUGACACAACCAAGAGCUUAUCUUCAAGAGAGAUUCGCUGUUCUACUGGAAAGGGAAUCCAUAGUAACGUAAGUUAU__CA_UGCU (((((...(((((((((((((((.....))))))..............(((......)))...)))))))))......((((.......))))......))))).............. (-19.17 = -17.87 + -1.30)

| Location | 5,533,642 – 5,533,760 |
|---|---|
| Length | 118 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 77.49 |
| Mean single sequence MFE | -33.88 |
| Consensus MFE | -22.34 |
| Energy contribution | -23.07 |
| Covariance contribution | 0.73 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.832082 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
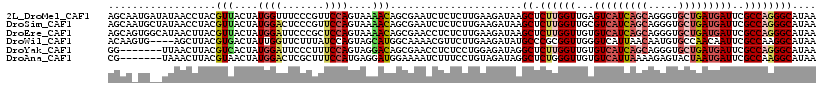
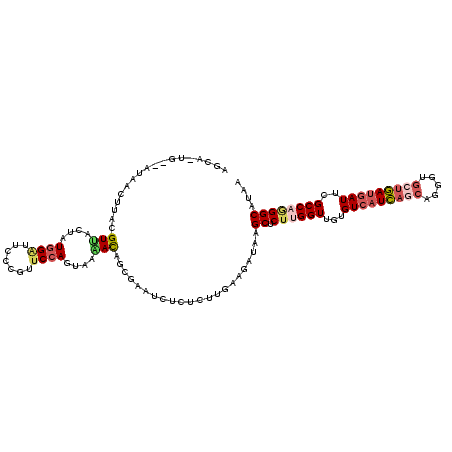
>2L_DroMel_CAF1 5533642 118 - 22407834 AGCAAUGAUAUAACCUACGUUACUAUGGUUUCCCGUUCCAGUAAAACAGCGAAUCUCUCUUGAAGAUAAGCUCUUGGUUGAGUCAUCAGCAGGGUGCUGAUGAUUCGCCAGGGCAUAA .((................(((((((((....))))...)))))...(((..(((((....).))))..)))((((((.(((((((((((.....))))))))))))))))))).... ( -37.10) >DroSim_CAF1 7670 118 - 1 AGCAAUGCUAUAACCUACGUUACUAUGGACUCCCGUUCCAGUAAAACAGCGAAUCUCUCUUGAAGAUAAGCUCUUGGUUGCGUCAUCAGCAGGGUGCUGAUGAUUCGCCAGGGCAUAA ....((((...........(((((((((....))))...)))))...(((..(((((....).))))..)))((((((.(.(((((((((.....))))))))).))))))))))).. ( -34.80) >DroEre_CAF1 7722 118 - 1 AGCAGUGGCAUAACUUACGUUACUAUGGAUUCCCGCUCCAGUAAAACAGCGAACCUCUCUUGAAGAUAAGCUCUUGGUUGUGUCAUCAGCAGGGUGCUGAUGAUUCGCCAGGGCAUAA .((((((((.........)))))).((((.......))))........))...................((.((((((...(((((((((.....)))))))))..)))))))).... ( -34.90) >DroWil_CAF1 7795 114 - 1 ACAAGUG----AGCUUACGUGACUAUUGGUUCUUUAUCCAGUAGCAUGGCAAAACGUUCUUGAAGAUAUGCCCGCGGUUGGGUCAUUAACAAUGUGCCAACAAUUCGCCAAGGCAUAA .((((..----......((((.(((((((........)))))))))))..........))))....((((((.((((((((..(((.....)))..)))))....)))...)))))). ( -32.17) >DroYak_CAF1 7740 111 - 1 GG-------UUAACUUACGUCACUAUGGAUUCCCUUUCCAGUAGGACAGCGAACCUCUCCUGGAGAUAGGCUCUUGGUUGUGUCAUCAGCAGGGUGCUGAUGAUUCGCCAGGGCAUAA ((-------((..((...(((.(((((((.......)))).))))))))..))))((((...))))...((.((((((...(((((((((.....)))))))))..)))))))).... ( -39.80) >DroAna_CAF1 7420 111 - 1 CG-------UAAACUUACGUAACUAUGGACUCGCUUUCCAUGAGGAUGGAAAAUCUUUCCUGUAGAUAGGCUCUGGGUUGUGUCAUUAAAAGAGUACUAAUGAUUCGCCAAGGCAUAA ((-------((....))))...(((((((.....(((((((....))))))).....))).))))....((.((.(((...(((((((.........)))))))..))).)))).... ( -24.50) >consensus AGCA_UG__AUAACUUACGUUACUAUGGAUUCCCGUUCCAGUAAAACAGCGAAUCUCUCUUGAAGAUAAGCUCUUGGUUGUGUCAUCAGCAGGGUGCUGAUGAUUCGCCAGGGCAUAA ..................(((....((((.......))))....)))......................((.((((((...(((((((((.....)))))))))..)))))))).... (-22.34 = -23.07 + 0.73)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:17:58 2006