
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 5,512,268 – 5,512,364 |
| Length | 96 |
| Max. P | 0.874322 |

| Location | 5,512,268 – 5,512,364 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 80.53 |
| Mean single sequence MFE | -23.52 |
| Consensus MFE | -13.77 |
| Energy contribution | -14.37 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.59 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.558174 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 5512268 96 + 22407834 GCAACAGUGACAGAGUUUUCUUUUUAAGUCACAAUCGGCGAAAGCCAAUCGCUUUGCUGCCUAAUAUUU-AUACCGUGUAAUAGAGGCAUUACU---AGA ((((..(((((((((....))))....)))))....(((....))).......))))(((((.....((-((.....))))...))))).....---... ( -24.80) >DroSec_CAF1 135992 96 + 1 GCAACAGUGACAGAGUUUUCUUUUUAAGUCACAAUCGGCGAAAGCCAAUCGCUUUGCUGCCUAAUAUUU-AUACCGUGUAAUAGAGGCAUUACU---AGA ((((..(((((((((....))))....)))))....(((....))).......))))(((((.....((-((.....))))...))))).....---... ( -24.80) >DroSim_CAF1 142983 96 + 1 GCAACAGUGACAGAGUUUUCUUUUUAAGUCACUAUCGGCGAAAGCCAAUCGCUUUGCUGCCUAAUAUUU-AUACCGUGCAAUAGAGGCAUUACU---AGA ((((.((((((((((....))))....))))))...(((....))).......))))(((((..(((((-((...))).)))).))))).....---... ( -23.70) >DroWil_CAF1 195719 91 + 1 ACAGCAGCAACAAAGCUUUCUUUUUGCU------GCGGCUCAUGGCGAUUUGUUUGCUGCCUAAUAUUUUCUACCUCAU---ACAAAUAUCAAUCGUUGA ...(((((((.((((....)))))))))------))(((....(((((.....))))))))..................---.................. ( -19.60) >DroYak_CAF1 141317 96 + 1 GCAACAGUGAAUGAGUUUUCUUUUUAAGUCGCAAUCGGCGAAAGCCAAUCGCUUUGCUGCCUAAUAUUU-AUACCGUGUAAUAGAGGCAUUGCU---AGA ((((.(((((..((.((........)).))......(((....)))..)))))))))(((((.....((-((.....))))...))))).....---... ( -24.70) >consensus GCAACAGUGACAGAGUUUUCUUUUUAAGUCACAAUCGGCGAAAGCCAAUCGCUUUGCUGCCUAAUAUUU_AUACCGUGUAAUAGAGGCAUUACU___AGA ((((.(((((.((((....)))).............(((....)))..)))))))))(((((......................)))))........... (-13.77 = -14.37 + 0.60)


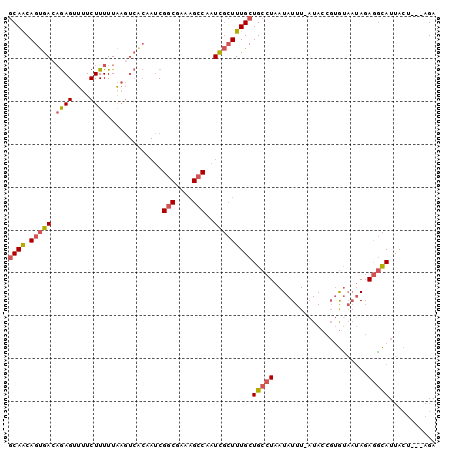
| Location | 5,512,268 – 5,512,364 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 80.53 |
| Mean single sequence MFE | -24.61 |
| Consensus MFE | -15.52 |
| Energy contribution | -15.88 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.36 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.874322 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 5512268 96 - 22407834 UCU---AGUAAUGCCUCUAUUACACGGUAU-AAAUAUUAGGCAGCAAAGCGAUUGGCUUUCGCCGAUUGUGACUUAAAAAGAAAACUCUGUCACUGUUGC .((---(((((((((..........)))))-...))))))((((((..(((((((((....)))))))))(((................)))..)))))) ( -26.49) >DroSec_CAF1 135992 96 - 1 UCU---AGUAAUGCCUCUAUUACACGGUAU-AAAUAUUAGGCAGCAAAGCGAUUGGCUUUCGCCGAUUGUGACUUAAAAAGAAAACUCUGUCACUGUUGC .((---(((((((((..........)))))-...))))))((((((..(((((((((....)))))))))(((................)))..)))))) ( -26.49) >DroSim_CAF1 142983 96 - 1 UCU---AGUAAUGCCUCUAUUGCACGGUAU-AAAUAUUAGGCAGCAAAGCGAUUGGCUUUCGCCGAUAGUGACUUAAAAAGAAAACUCUGUCACUGUUGC .((---(((((((((..........)))))-...))))))(((((......((((((....))))))((((((................))))))))))) ( -25.69) >DroWil_CAF1 195719 91 - 1 UCAACGAUUGAUAUUUGU---AUGAGGUAGAAAAUAUUAGGCAGCAAACAAAUCGCCAUGAGCCGC------AGCAAAAAGAAAGCUUUGUUGCUGCUGU .......(((((((((..---..........)))))))))((((((........((.....)).((------((((((........)))))))))))))) ( -20.20) >DroYak_CAF1 141317 96 - 1 UCU---AGCAAUGCCUCUAUUACACGGUAU-AAAUAUUAGGCAGCAAAGCGAUUGGCUUUCGCCGAUUGCGACUUAAAAAGAAAACUCAUUCACUGUUGC ...---....(((((..........)))))-.........((((((..(((((((((....)))))))))..........(((......)))..)))))) ( -24.20) >consensus UCU___AGUAAUGCCUCUAUUACACGGUAU_AAAUAUUAGGCAGCAAAGCGAUUGGCUUUCGCCGAUUGUGACUUAAAAAGAAAACUCUGUCACUGUUGC ...........((((..........))))...........((((((..(((((((((....))))))))).........(((....))).....)))))) (-15.52 = -15.88 + 0.36)


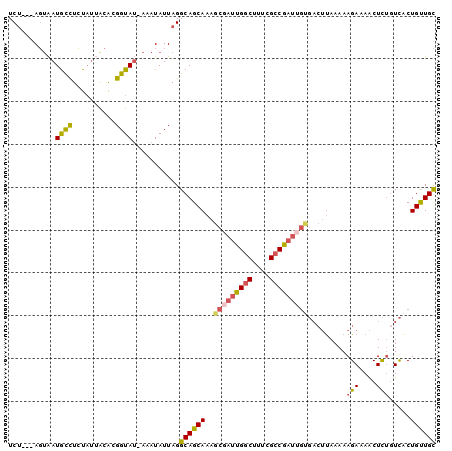
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:17:50 2006