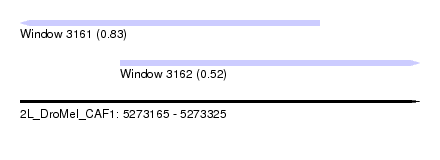
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 5,273,165 – 5,273,325 |
| Length | 160 |
| Max. P | 0.830835 |

| Location | 5,273,165 – 5,273,285 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.44 |
| Mean single sequence MFE | -42.73 |
| Consensus MFE | -35.71 |
| Energy contribution | -35.85 |
| Covariance contribution | 0.14 |
| Combinations/Pair | 1.31 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.830835 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 5273165 120 - 22407834 GCCUUAAGUUCUUCGAGCUCUCCCUCAAGGAGCAAAUCAAGGAUCUGAGUCUGCAGCGGGACGGUCUGGUCAUGGAACUGCAGCAGUUGCAGGAGGCGCGACCCGUGCUCGAGAAGGCCU (((((......(((((((..(((.....)))(((..(((......)))...))).(((((.((((((..((.((.(((((...))))).)))))))).)).))))))))))))))))).. ( -40.40) >DroVir_CAF1 3214 120 - 1 GUCUUAAGUUCUUUGAGCUUUCACUCAAGGAGCAGAUAAAGGAUCUGAGUCUGCAACGUGACGGUUUGGUCAUGGAAUUGCAACAGUUGCAGGAAGCACGUCCUGUUCUUCAGAAAGCAU .(((...((((((((((......))))))))))(((...(((((.((..((((((((.((.((((((.......))))))))...))))))))...)).)))))..)))..)))...... ( -39.60) >DroGri_CAF1 524 120 - 1 GCCUCAAAUUCUUUGAGCUCUCACUCAGAGAGCAGAUCAAGGAUUUGAGCCUGCAACGUGAUGGUUUAGUAAUGGAACUGCAACAGUUGCAGGAAGCGCGUCCAGUUCUCGAGAAGGCCU (((((((((.((((((((((((.....))))))...)))))))))))))((((((((.((.((((((.......))))))))...))))))))..))((.((..(....)..))..)).. ( -41.90) >DroWil_CAF1 2974 120 - 1 GCCUUAAAUUCUUUGAAUUGUCACUAAAGGAACACAUCAAAGAUUUGAGCCUGCAACGUGACGGUUUGGUCAUGGAACUGCAACAGUUGCAGGAGGCGCGUCCAGUACUUGAAAAGGCCU (((((....(((((((..(((..(....)..)))..)))))))((..((((((((((.((.((((((.......))))))))...)))))))).((.....))....))..))))))).. ( -38.20) >DroMoj_CAF1 498 120 - 1 GUCUCAAGUUCUUUGAGCUUUCGCUCAAGGAGCAGAUCAAGGAUUUGAGCCUGCAACGUGACGGUUUAGUCAUGGAACUGCAACAGUUGCAGGAAGCACGUCCCGUUCUCGAGAAGGCAU .((((..(((((((((((....)))))))))))(((....((((.((..((((((((.((.((((((.......))))))))...))))))))...)).))))...))).))))...... ( -49.90) >DroAna_CAF1 28907 120 - 1 GCCUCAAGUUCUUCGAGCUGUCGCUCAAGGAGCAGAUCAAGGACUUGAGCCUGCAACGCGACGGUUUGGUCAUGGAACUGCAGCAGUUGCAGGAGGCGCGACCCGUAUUAGAGAAGGCCU ..(((((((((((.((.((((..(....)..)))).)))))))))))))((((((((((..((((((.......))))))..)).))))))))((((.(..(.(......).)..))))) ( -46.40) >consensus GCCUCAAGUUCUUUGAGCUCUCACUCAAGGAGCAGAUCAAGGAUUUGAGCCUGCAACGUGACGGUUUGGUCAUGGAACUGCAACAGUUGCAGGAAGCGCGUCCCGUUCUCGAGAAGGCCU ((((...((((((((((......)))))))))).......((((.((..((((((((((..((((((.......))))))..)).))))))))...)).))))...........)))).. (-35.71 = -35.85 + 0.14)
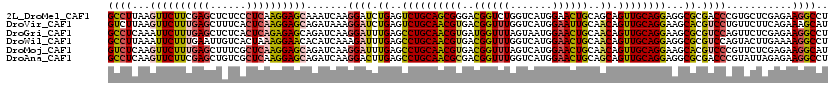

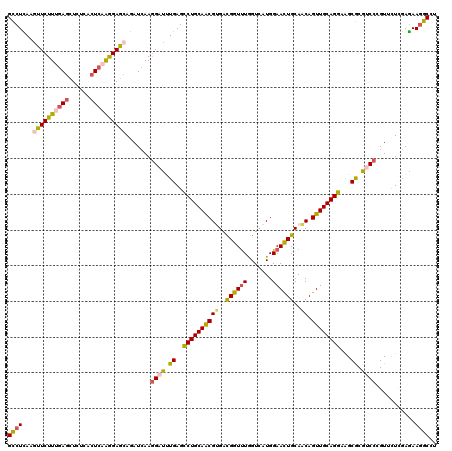
| Location | 5,273,205 – 5,273,325 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.33 |
| Mean single sequence MFE | -32.93 |
| Consensus MFE | -22.15 |
| Energy contribution | -21.18 |
| Covariance contribution | -0.96 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.67 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.516645 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 5273205 120 + 22407834 CAGUUCCAUGACCAGACCGUCCCGCUGCAGACUCAGAUCCUUGAUUUGCUCCUUGAGGGAGAGCUCGAAGAACUUAAGGCGCUUCAGUUCCUCCUUCAGUGCCGACGGACUGCUGCACUU ........((..(((.(((((.(((((((((.((((....))))))))).....(((((((((((.((((..(.....)..)))))))).)))))))))))..))))).)))...))... ( -38.60) >DroVir_CAF1 3254 120 + 1 CAAUUCCAUGACCAAACCGUCACGUUGCAGACUCAGAUCCUUUAUCUGCUCCUUGAGUGAAAGCUCAAAGAACUUAAGACGUUUCAGUUCCUCCUUCAGAGCACUUGGACUACUGCACUU ........((((......))))...(((((....((.(((......(((((.((((((....)))))).(((((.(((...))).)))))........)))))...))))).)))))... ( -26.90) >DroGri_CAF1 564 120 + 1 CAGUUCCAUUACUAAACCAUCACGUUGCAGGCUCAAAUCCUUGAUCUGCUCUCUGAGUGAGAGCUCAAAGAAUUUGAGGCGCUUCAGUUCUUCCUUUAAAAGAUUCGGACUACUGCACUU (((((((................(..((.(.(((((((.(((.....((((((.....))))))...))).))))))).)))..)...((((.......))))...)))..))))..... ( -29.60) >DroWil_CAF1 3014 120 + 1 CAGUUCCAUGACCAAACCGUCACGUUGCAGGCUCAAAUCUUUGAUGUGUUCCUUUAGUGACAAUUCAAAGAAUUUAAGGCGUUUCAGUUCCUCCUUCAGUAUGCUUGGGCUGCUGCAUUU ........((((......)))).((.((((.(((((.(((((((..(((..(....)..)))..)))))))(((.((((.(.........).)))).)))....))))))))).)).... ( -27.70) >DroMoj_CAF1 538 120 + 1 CAGUUCCAUGACUAAACCGUCACGUUGCAGGCUCAAAUCCUUGAUCUGCUCCUUGAGCGAAAGCUCAAAGAACUUGAGACGCUUUAAUUCCUCCUUAAGGACACUUGGGCUAUUACAUUU .(((.(((.(..((((.((((.....(((((.((((....)))))))))((.((((((....)))))).))......)))).))))..((((.....))))..).))))))......... ( -33.80) >DroAna_CAF1 28947 120 + 1 CAGUUCCAUGACCAAACCGUCGCGUUGCAGGCUCAAGUCCUUGAUCUGCUCCUUGAGCGACAGCUCGAAGAACUUGAGGCGCUUCAGCUCCUCCUUGAGGACCGAGGGACUGCUGCACUU (((((((..(((......)))((((((((((.((((....)))))))))((.((((((....)))))).))......))))).((.(.(((((...)))))).)))))))))........ ( -41.00) >consensus CAGUUCCAUGACCAAACCGUCACGUUGCAGGCUCAAAUCCUUGAUCUGCUCCUUGAGUGAAAGCUCAAAGAACUUAAGGCGCUUCAGUUCCUCCUUCAGAACACUUGGACUACUGCACUU .................((((.....(((((.((((....)))))))))((.((((((....)))))).))......))))...((((...(((............)))..))))..... (-22.15 = -21.18 + -0.96)

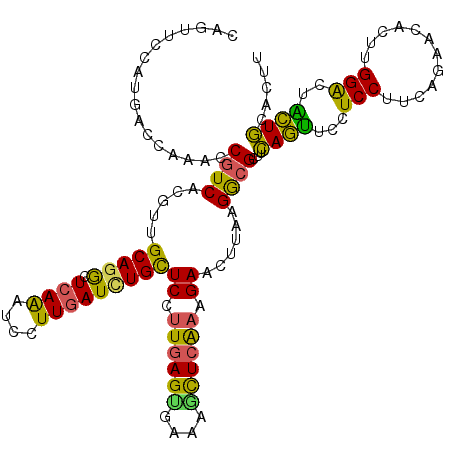

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:15:51 2006