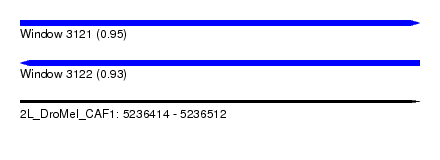

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 5,236,414 – 5,236,512 |
| Length | 98 |
| Max. P | 0.948284 |

| Location | 5,236,414 – 5,236,512 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 94.06 |
| Mean single sequence MFE | -36.20 |
| Consensus MFE | -27.40 |
| Energy contribution | -27.32 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.09 |
| Mean z-score | -3.02 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.38 |
| SVM RNA-class probability | 0.948284 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 5236414 98 + 22407834 GGGCCAAAGGCUGAUGAUUUUUAUACGGCGGCAACAACUGCAAUGCAACUAAGUUGUUGUUGCCCCAACCGAUCG-UUCUUGGUU---GUUUUUGUUGGCCU (((((((...................(((((((((((((............))))))))))))).(((((((...-...))))))---)......))))))) ( -36.90) >DroSec_CAF1 13664 98 + 1 GGGCCAAAGGCUGAUGAUUUUUAUACGGCGGCGGCCACUGCAGUGCAACUCAGUUGUUGUUGCCCCAACCGAUCG-UUCUUGGUU---GUUUUUGUUGGCCU (((((((...................(((((((((.((((.((.....)))))).))))))))).(((((((...-...))))))---)......))))))) ( -36.40) >DroSim_CAF1 14546 98 + 1 GGGCCAAAGGCUGAUGAUUUUUAUACGGCGGCGGCAACUGCAAUGCAACUUAGUUGUUGUUGCCCCAACCGAUCG-UUCUUGGUU---GUUUUUGUUGGCCU (((((((..((((((((...)))).))))((.((((((.(((((........))))).))))))))((((((...-...))))))---.......))))))) ( -36.70) >DroEre_CAF1 6317 99 + 1 GGGCCAAAGGCUGAUGAUUUUUAUACGGCGGCGG---CUGCAAUGCAACUUAGUUGUUGUUGCCCCAACCGAUCGGUUCUUGGUUGUUGUUUUUGUUGGCCU (((((((..((((((((...)))).))))(((((---(.(((((........))))).)))))).(((((((.......))))))).........))))))) ( -34.20) >DroYak_CAF1 15648 95 + 1 GGGCCAAAGGCUGAUGAUUUUUAUACGGCGGCGG---CUGCAAUGCAACUUAGUUGUUGUUGCCACAACCGAUCG-UUCUUGGUU---GUUUUUGUUGGCCU (((((((..((((((((...)))).))))(((((---(.(((((........))))).))))))((((((((...-...))))))---)).....))))))) ( -36.80) >consensus GGGCCAAAGGCUGAUGAUUUUUAUACGGCGGCGGC_ACUGCAAUGCAACUUAGUUGUUGUUGCCCCAACCGAUCG_UUCUUGGUU___GUUUUUGUUGGCCU (((((((..((((((((...)))).))))((((((....(((((........))))).))))))..((((((.......))))))..........))))))) (-27.40 = -27.32 + -0.08)

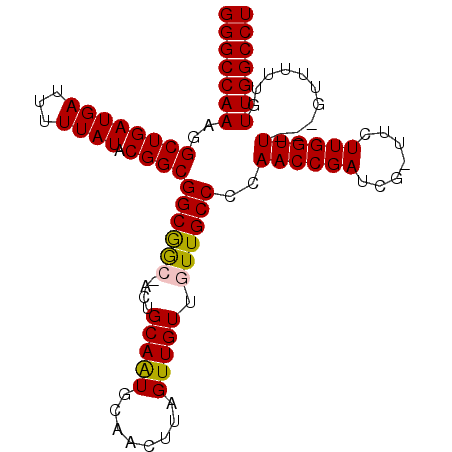
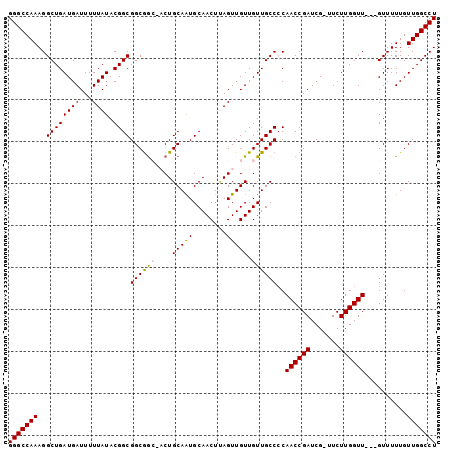
| Location | 5,236,414 – 5,236,512 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 94.06 |
| Mean single sequence MFE | -26.66 |
| Consensus MFE | -20.96 |
| Energy contribution | -21.00 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.929672 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 5236414 98 - 22407834 AGGCCAACAAAAAC---AACCAAGAA-CGAUCGGUUGGGGCAACAACAACUUAGUUGCAUUGCAGUUGUUGCCGCCGUAUAAAAAUCAUCAGCCUUUGGCCC .((((((......(---((((.....-.....))))).(((..((((((((..((......))))))))))..)))...................)))))). ( -28.90) >DroSec_CAF1 13664 98 - 1 AGGCCAACAAAAAC---AACCAAGAA-CGAUCGGUUGGGGCAACAACAACUGAGUUGCACUGCAGUGGCCGCCGCCGUAUAAAAAUCAUCAGCCUUUGGCCC .((((((......(---((((.....-.....)))))((((.......((((((.....)).))))(((....)))...............)))))))))). ( -28.60) >DroSim_CAF1 14546 98 - 1 AGGCCAACAAAAAC---AACCAAGAA-CGAUCGGUUGGGGCAACAACAACUAAGUUGCAUUGCAGUUGCCGCCGCCGUAUAAAAAUCAUCAGCCUUUGGCCC .((((((.......---......((.-...))(((.(.(((((((((......)))((...)).)))))).).)))...................)))))). ( -25.30) >DroEre_CAF1 6317 99 - 1 AGGCCAACAAAAACAACAACCAAGAACCGAUCGGUUGGGGCAACAACAACUAAGUUGCAUUGCAG---CCGCCGCCGUAUAAAAAUCAUCAGCCUUUGGCCC .((((((.........(((((...........)))))((((..((((......))))...(((.(---(....)).)))............)))))))))). ( -25.20) >DroYak_CAF1 15648 95 - 1 AGGCCAACAAAAAC---AACCAAGAA-CGAUCGGUUGUGGCAACAACAACUAAGUUGCAUUGCAG---CCGCCGCCGUAUAAAAAUCAUCAGCCUUUGGCCC .((((((.....((---((((.....-.....))))))(((..((((......))))...(((.(---(....)).)))............))).)))))). ( -25.30) >consensus AGGCCAACAAAAAC___AACCAAGAA_CGAUCGGUUGGGGCAACAACAACUAAGUUGCAUUGCAGU_GCCGCCGCCGUAUAAAAAUCAUCAGCCUUUGGCCC .((((((................((.....))((((((.(((((.........)))))((.((.(......).)).))..........)))))).)))))). (-20.96 = -21.00 + 0.04)

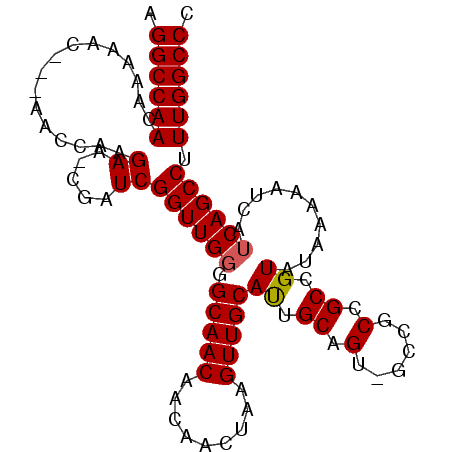
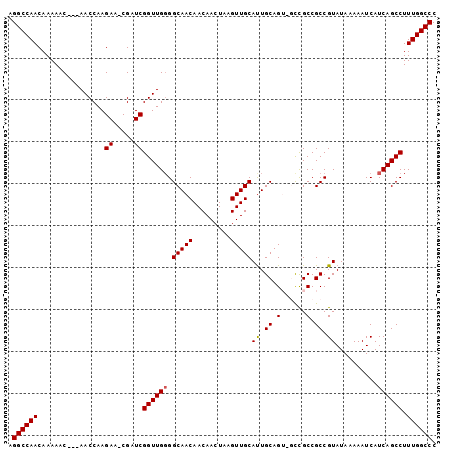
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:15:13 2006