
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 5,186,067 – 5,186,160 |
| Length | 93 |
| Max. P | 0.714629 |

| Location | 5,186,067 – 5,186,160 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 80.88 |
| Mean single sequence MFE | -39.38 |
| Consensus MFE | -21.90 |
| Energy contribution | -23.57 |
| Covariance contribution | 1.67 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.56 |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.714629 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 5186067 93 + 22407834 GUUGC---UGUUGUGGCCAGGUUCCGACCAACGGACGCUGCUGCGGCAACACAUUGCAACAG---GCAGCA---GCAGCAGAUCAGGGAAAUUGCCCGCCGU (((((---((((((...(((((.(((.....))))).)))(((..((((....))))..)))---))))))---)))))......(((......)))..... ( -36.30) >DroVir_CAF1 103015 90 + 1 GUUGUUGUUGUUGCGGCCCUGCCAGCAGCAGCACAUGCUG---------CACAUUGCUGCCA---CCAGCAACAGAAACGCACCCUGGUAAUGGCCCGCAGU ..........((((((((.((((((..(((((....))))---------)...((((((...---.))))))............))))))..)).)))))). ( -34.20) >DroSec_CAF1 63290 96 + 1 GUUGC---UGUUGUGGCCAGGUUUCGACCAACGGAUGCUGCUGCGGCAACACAUUGCAACAG---GCAGCAGCAGCAGCAGAUCCGGGAAAUUGCCCGCCGU (((((---((((((.(((.(((....)))......(((...((.(....).))..)))...)---)).))))))))))).....((((......)))).... ( -38.60) >DroSim_CAF1 36294 96 + 1 GUUGC---UGUUGUGGCCAGGUUCCGACCAACGGAUGCUGCUGCGGCAACACAUUGCAACAG---GCAGCAGCAGCAGCAGAUCCGGGAAAUUGCCCGCCGU (((((---((((((.(((((..((((.....))))..)))(((..((((....))))..)))---)).))))))))))).....((((......)))).... ( -40.40) >DroEre_CAF1 65615 99 + 1 GUUGC---UGUUGUGGCCAGGCUCCGACCAACGGAUGCUGCUGCGGCAACACCUUGCAACACAUUGCGGCAGCAGCAACAGAUCCGGGAAACUGCCCGCCGU (((((---((((((.((..(((((((.....)))).)))..((..((((....))))..))....)).))))))))))).....((((......)))).... ( -40.20) >DroYak_CAF1 65066 96 + 1 GUUGC---UGUUGUGGCCAGGUUCCGACCAACGGAUGCUGCUGCAGCAACGCAUUGCAGCAG---GCAGCAGCAGCAACAGAUCCGGGAAAUUGCCCGCCGU (((((---((((((.(((....((((.....))))(((((((((......)))..)))))))---)).))))))))))).....((((......)))).... ( -46.60) >consensus GUUGC___UGUUGUGGCCAGGUUCCGACCAACGGAUGCUGCUGCGGCAACACAUUGCAACAG___GCAGCAGCAGCAACAGAUCCGGGAAAUUGCCCGCCGU (((((...((((((.((..(((((((.....)))).)))...)).))))))....))))).....((.((....)).........(((......)))))... (-21.90 = -23.57 + 1.67)



| Location | 5,186,067 – 5,186,160 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 80.88 |
| Mean single sequence MFE | -38.55 |
| Consensus MFE | -22.49 |
| Energy contribution | -23.94 |
| Covariance contribution | 1.45 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.557232 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 5186067 93 - 22407834 ACGGCGGGCAAUUUCCCUGAUCUGCUGC---UGCUGC---CUGUUGCAAUGUGUUGCCGCAGCAGCGUCCGUUGGUCGGAACCUGGCCACAACA---GCAAC .(((.(((.....))))))...(((((.---.(((((---.(((.((((....)))).))))))))....((.((((((...))))))))..))---))).. ( -33.90) >DroVir_CAF1 103015 90 - 1 ACUGCGGGCCAUUACCAGGGUGCGUUUCUGUUGCUGG---UGGCAGCAAUGUG---------CAGCAUGUGCUGCUGCUGGCAGGGCCGCAACAACAACAAC ..(((((.((.....(((((.....))))).(((..(---..(((((((((..---------...))).))))))..)..))))).)))))........... ( -41.50) >DroSec_CAF1 63290 96 - 1 ACGGCGGGCAAUUUCCCGGAUCUGCUGCUGCUGCUGC---CUGUUGCAAUGUGUUGCCGCAGCAGCAUCCGUUGGUCGAAACCUGGCCACAACA---GCAAC ....((((......))))....(((((.(((((((((---.....((((....)))).)))))))))...((.(((((.....)))))))..))---))).. ( -36.40) >DroSim_CAF1 36294 96 - 1 ACGGCGGGCAAUUUCCCGGAUCUGCUGCUGCUGCUGC---CUGUUGCAAUGUGUUGCCGCAGCAGCAUCCGUUGGUCGGAACCUGGCCACAACA---GCAAC ....((((......))))....(((((.(((((((((---.....((((....)))).)))))))))...((.((((((...))))))))..))---))).. ( -37.90) >DroEre_CAF1 65615 99 - 1 ACGGCGGGCAGUUUCCCGGAUCUGUUGCUGCUGCCGCAAUGUGUUGCAAGGUGUUGCCGCAGCAGCAUCCGUUGGUCGGAGCCUGGCCACAACA---GCAAC ..(((((((................(((((((((.((((..(.(....).)..)))).)))))))))((((.....)))))))).)))......---..... ( -41.20) >DroYak_CAF1 65066 96 - 1 ACGGCGGGCAAUUUCCCGGAUCUGUUGCUGCUGCUGC---CUGCUGCAAUGCGUUGCUGCAGCAGCAUCCGUUGGUCGGAACCUGGCCACAACA---GCAAC ....((((......)))).....(((((((.((((((---.(((.((((....)))).)))))))))...((.((((((...))))))))..))---))))) ( -40.40) >consensus ACGGCGGGCAAUUUCCCGGAUCUGCUGCUGCUGCUGC___CUGUUGCAAUGUGUUGCCGCAGCAGCAUCCGUUGGUCGGAACCUGGCCACAACA___GCAAC ...(((((......)))(((((((((((.(((((.((.....)).))).......)).))))))).))))((.((((((...)))))))).......))... (-22.49 = -23.94 + 1.45)

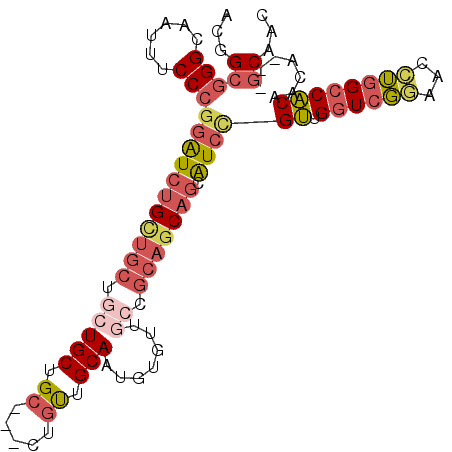

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:13:48 2006