
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 4,904,320 – 4,904,417 |
| Length | 97 |
| Max. P | 0.939915 |

| Location | 4,904,320 – 4,904,417 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 83.06 |
| Mean single sequence MFE | -38.92 |
| Consensus MFE | -28.52 |
| Energy contribution | -29.55 |
| Covariance contribution | 1.03 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.67 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.848837 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 4904320 97 + 22407834 UAAGAGCAGGAGGUGGCAGCUUAAAGCCACUUGGACCACCCAAAUUAAGGGAUCCUCCUGUAAAGACAGGAGGCACUGUUGUGGUGGU---UGUGGUGCU ....((((..(((((((........))))))).((((((((((....((....((((((((....))))))))..)).))).))))))---)....)))) ( -41.30) >DroSec_CAF1 102 97 + 1 UCAGAGCAGGUGGUGGCAAUUUGAAGCCACUUGGACCACCCAAAUUAAGGGAUCCUCCUGUGAAGACAGGAGGCACUGUUGUGGUGGU---UGUGGUGCU ......((((((((..(.....)..))))))))((((((((((....((....((((((((....))))))))..)).))).))))))---)........ ( -40.70) >DroSim_CAF1 102 97 + 1 UGAGAGCAGGUGGUGGCAGUUUGAAGCCACUUGGUCCACCCAAAUUAAGGGAUCCUCCUGUGAAGAUAGGAGGCACUGUUGUGGUGGU---UGUGGUGCU ...((.((((((((..(.....)..)))))))).))(((((((..........((((((((....))))))))((((.....)))).)---)).)))).. ( -36.70) >DroEre_CAF1 101 97 + 1 UGAGAGCAGGUGGAGGCAGCCUAAAGCCACUUGGACCACCCAAUUUGAGGGAUCCUCCUGCGAAGGCAGGAGGCACUGUUGUGGUGGA---UGUGGUGCU ....(((((((.......))))...(((((.....(((((((((....((...((((((((....))))))))..)))))).))))).---.)))))))) ( -41.70) >DroYak_CAF1 296 97 + 1 UGAGAGCAGGUGGUGGCAGCUUAAAACCACUUGGACCACCCAAUUUGAGGGAUCCUCCUGUGAAGACAGGAGACGCUGUUGUGGUGGA---UGUGGUGCU ......((((((((...........)))))))).((((((((...(((.((...(((((((....)))))))...)).)))...))).---.)))))... ( -34.20) >DroPer_CAF1 105 100 + 1 CGAGGGCUGGCGGUGGCAGCUUGAACCCACUGGGGCCACCUAAAUUGAGAGGACCACCUGCAUAGGGCGGAGGAGCCGCAGUGGUGCUACUCGUACUGCU ....(((..(((((((((........((((((.((((.(((........))).((.(((.....))).))..).))).)))))))))))).)))...))) ( -38.90) >consensus UGAGAGCAGGUGGUGGCAGCUUAAAGCCACUUGGACCACCCAAAUUAAGGGAUCCUCCUGUGAAGACAGGAGGCACUGUUGUGGUGGU___UGUGGUGCU ....((((...((((((........))))))....((((((((.....((...((((((((....))))))))..)).))).))))).........)))) (-28.52 = -29.55 + 1.03)


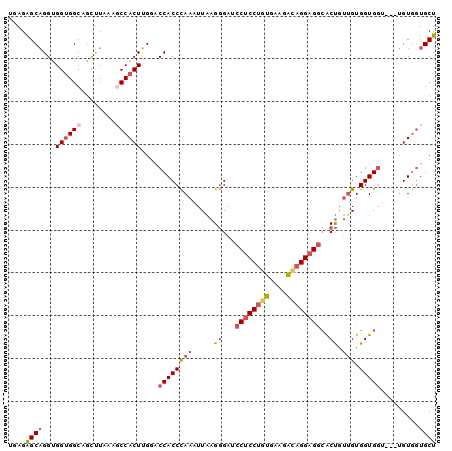
| Location | 4,904,320 – 4,904,417 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 83.06 |
| Mean single sequence MFE | -31.85 |
| Consensus MFE | -25.27 |
| Energy contribution | -25.80 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.20 |
| Mean z-score | -3.13 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.28 |
| SVM RNA-class probability | 0.939915 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 4904320 97 - 22407834 AGCACCACA---ACCACCACAACAGUGCCUCCUGUCUUUACAGGAGGAUCCCUUAAUUUGGGUGGUCCAAGUGGCUUUAAGCUGCCACCUCCUGCUCUUA ((((.....---((((((.(((.((..((((((((....))))))))....))....)))))))))....(((((........)))))....)))).... ( -34.00) >DroSec_CAF1 102 97 - 1 AGCACCACA---ACCACCACAACAGUGCCUCCUGUCUUCACAGGAGGAUCCCUUAAUUUGGGUGGUCCAAGUGGCUUCAAAUUGCCACCACCUGCUCUGA ((((.....---((((((.(((.((..((((((((....))))))))....))....)))))))))....(((((........)))))....)))).... ( -34.10) >DroSim_CAF1 102 97 - 1 AGCACCACA---ACCACCACAACAGUGCCUCCUAUCUUCACAGGAGGAUCCCUUAAUUUGGGUGGACCAAGUGGCUUCAAACUGCCACCACCUGCUCUCA ((((.....---.(((((.(((.((..((((((........))))))....))....)))))))).....(((((........)))))....)))).... ( -29.50) >DroEre_CAF1 101 97 - 1 AGCACCACA---UCCACCACAACAGUGCCUCCUGCCUUCGCAGGAGGAUCCCUCAAAUUGGGUGGUCCAAGUGGCUUUAGGCUGCCUCCACCUGCUCUCA (((......---.....(((....)))((((((((....))))))))............((((((.....(..((.....))..)..))))))))).... ( -32.40) >DroYak_CAF1 296 97 - 1 AGCACCACA---UCCACCACAACAGCGUCUCCUGUCUUCACAGGAGGAUCCCUCAAAUUGGGUGGUCCAAGUGGUUUUAAGCUGCCACCACCUGCUCUCA ((((.....---.(((((.(((.((..((((((((....))))))))....))....)))))))).....(((((........)))))....)))).... ( -28.10) >DroPer_CAF1 105 100 - 1 AGCAGUACGAGUAGCACCACUGCGGCUCCUCCGCCCUAUGCAGGUGGUCCUCUCAAUUUAGGUGGCCCCAGUGGGUUCAAGCUGCCACCGCCAGCCCUCG .((.....(((.((.(((((((((((......)))....))).))))).)))))......(((((((((...)))........))))))))......... ( -33.00) >consensus AGCACCACA___ACCACCACAACAGUGCCUCCUGUCUUCACAGGAGGAUCCCUCAAUUUGGGUGGUCCAAGUGGCUUCAAGCUGCCACCACCUGCUCUCA .((..........(((((.(((.((..((((((((....))))))))....))....)))))))).....(((((........))))).....))..... (-25.27 = -25.80 + 0.53)


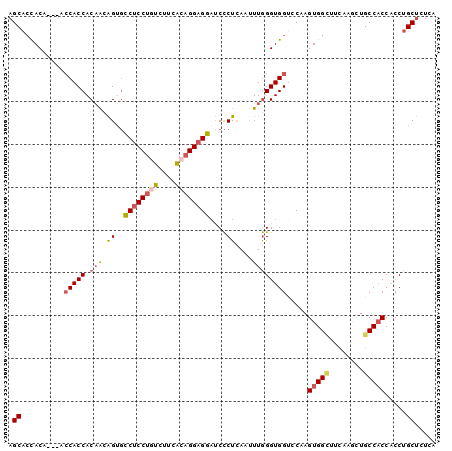
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:11:23 2006