
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 4,891,298 – 4,891,503 |
| Length | 205 |
| Max. P | 0.996807 |

| Location | 4,891,298 – 4,891,389 |
|---|---|
| Length | 91 |
| Sequences | 3 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 70.35 |
| Mean single sequence MFE | -13.13 |
| Consensus MFE | -8.29 |
| Energy contribution | -8.07 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.691190 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
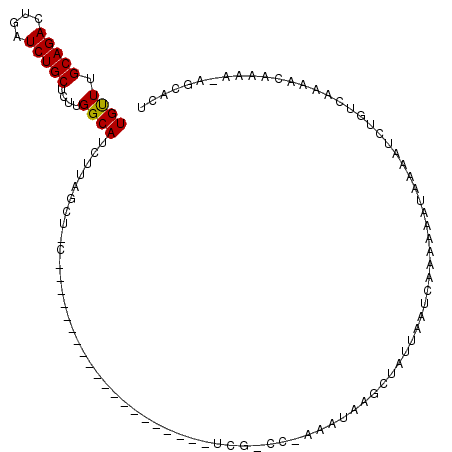
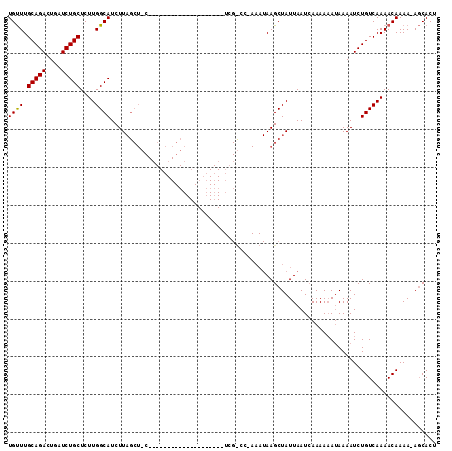
>2L_DroMel_CAF1 4891298 91 + 22407834 UGCUUGCAGACUGAUCUGCUCUUGGCAUCUUAGCUUC--------------------UCGUCC-AAAUAAGCUAUUAAUAAAAA-AAAAAAUCUGUCAAAACAAAAAACCACU ((((.(((((....)))))....))))...((((((.--------------------......-....))))))..........-............................ ( -13.10) >DroSec_CAF1 1228 75 + 1 UGUUUGCAGACUCAUCUGCUCUUGGCAUCUU-------------------------------------AAGCUAUUAAUCAAAAAAUAAAAUCUGUCAAAACAAAA-AGCACU ((((((((((....)))))...((((.....-------------------------------------..))))........................)))))...-...... ( -10.40) >DroSim_CAF1 1219 112 + 1 UGUUUGCAGAAUGCUCUGCUCUUGGCAUCUUAGCUGCUCGAAAAGCAAACAAAUCUCUCGCCCAAAAUAAGCUAUUAAUCAAAAAAUAAAAUCUGUCAAAACAAAA-AGCACU ((((((((((....)))))..((((((.....(((........))).............((.........)).....................))))))......)-)))).. ( -15.90) >consensus UGUUUGCAGACUGAUCUGCUCUUGGCAUCUUAGCU_C____________________UCG_CC_AAAUAAGCUAUUAAUCAAAAAAUAAAAUCUGUCAAAACAAAA_AGCACU ((((.(((((....)))))....))))...................................................................................... ( -8.29 = -8.07 + -0.22)

| Location | 4,891,389 – 4,891,503 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.81 |
| Mean single sequence MFE | -35.74 |
| Consensus MFE | -30.10 |
| Energy contribution | -31.58 |
| Covariance contribution | 1.48 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.91 |
| SVM RNA-class probability | 0.982414 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
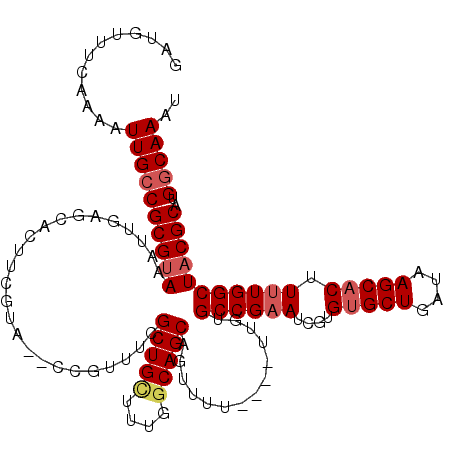
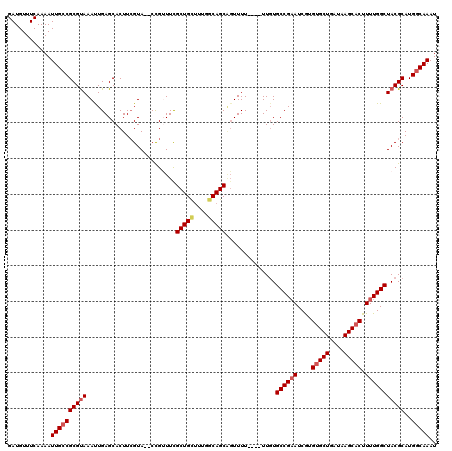
>2L_DroMel_CAF1 4891389 114 + 22407834 GAUGUUUCAAAAUUGCCGCGUAAAUUGAGCACUUGGUA--CCGUUUUGCUGUAUCGGCAGCAGUUUU----UUGUGCCGAAUCGUGUGCUGAUAAGCACUUUUGGCUACGCAUGGCAAAU ............((((((((((....((....((((((--(....(((((((....)))))))....----..))))))).))..(((((....))))).......)))))..))))).. ( -35.90) >DroSec_CAF1 1303 114 + 1 GAUGUUUCAAAAUUGCCGCGUAAAUUGAGCACUUCGUA--CCGUUUCGCUGCUUUGGCAGCCGUUUU----UUGUGCCGAAUCGUGUGCUGAUAAGCUCUUUUGGCUACGCAUGGCAAAU ............((((((((((((..((((...(((((--(((.((((((((....))))).((...----....)).))).)).)))).))...))))..))...)))))..))))).. ( -34.40) >DroSim_CAF1 1331 118 + 1 GAUGUUUCAAAAUUGCCGCGCAAAUUGAGCACUUCGUA--CCGUUUCGCUGCUUCGGCAGCAGUUUUUUUUUUUUGCCGAAUCGUGUGCUGAUAAGCACUUUUGGCUACGCAUGGCAAAU ............((((((.(((((..((((((......--..))...(((((....))))).))))..)))....((((((....(((((....))))).))))))...)).)))))).. ( -38.60) >DroEre_CAF1 1403 114 + 1 GAUGUUUCAAAAUUGCCGCGUAAACUGAGCACUUCGUA--CUGAUUCGCUGCUUUGGCAGCGUUUUU----UUGUGCCGAAUCAAGUGCUGAUAAGCACUUUUGGCUACGCAUGCCAAAU ..((.(((.........((.........)).....(((--(.((..((((((....))))))..)).----..)))).))).))((((((....))))))((((((.......)))))). ( -33.90) >DroYak_CAF1 1380 116 + 1 GAUGUUUCAAAAUUGCCGCGUAAACUGAGCACUUCGUACACCGAUUCGCUGCUUUGCCAGCAGUUUU----UUGUGCCGUAUCGAGUGCUGAUAAGCACUUUUGGCUACGCAUGGCAAAU ............((((((((((....(.((((.(((.....)))...((((((.....))))))...----..)))))((...(((((((....)))))))...)))))))..))))).. ( -35.90) >consensus GAUGUUUCAAAAUUGCCGCGUAAAUUGAGCACUUCGUA__CCGUUUCGCUGCUUUGGCAGCAGUUUU____UUGUGCCGAAUCGUGUGCUGAUAAGCACUUUUGGCUACGCAUGGCAAAU ............((((((((((.........................(((((....)))))..............((((((....(((((....))))).)))))))))))..))))).. (-30.10 = -31.58 + 1.48)

| Location | 4,891,389 – 4,891,503 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.81 |
| Mean single sequence MFE | -33.57 |
| Consensus MFE | -27.92 |
| Energy contribution | -29.08 |
| Covariance contribution | 1.16 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.58 |
| Structure conservation index | 0.83 |
| SVM decision value | 2.75 |
| SVM RNA-class probability | 0.996807 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
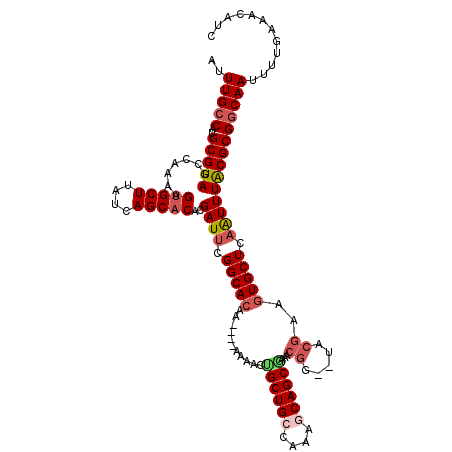
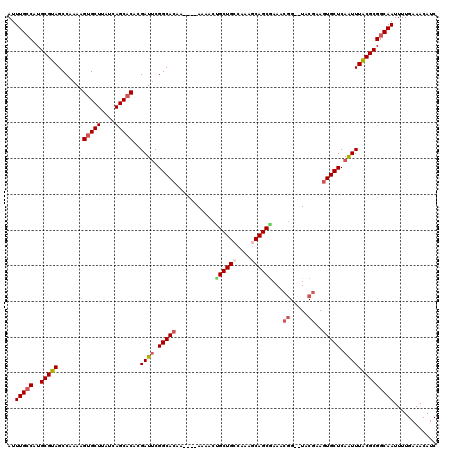
>2L_DroMel_CAF1 4891389 114 - 22407834 AUUUGCCAUGCGUAGCCAAAAGUGCUUAUCAGCACACGAUUCGGCACAA----AAAACUGCUGCCGAUACAGCAAAACGG--UACCAAGUGCUCAAUUUACGCGGCAAUUUUGAAACAUC ..(((((..(((((.......(((((....)))))..((((.(((((..----.....(((((......)))))....(.--...)..))))).))))))))))))))............ ( -32.30) >DroSec_CAF1 1303 114 - 1 AUUUGCCAUGCGUAGCCAAAAGAGCUUAUCAGCACACGAUUCGGCACAA----AAAACGGCUGCCAAAGCAGCGAAACGG--UACGAAGUGCUCAAUUUACGCGGCAAUUUUGAAACAUC ..(((((..(((((.........(((....)))....((((.(((((..----......(((((....)))))....((.--..))..))))).))))))))))))))............ ( -29.30) >DroSim_CAF1 1331 118 - 1 AUUUGCCAUGCGUAGCCAAAAGUGCUUAUCAGCACACGAUUCGGCAAAAAAAAAAAACUGCUGCCGAAGCAGCGAAACGG--UACGAAGUGCUCAAUUUGCGCGGCAAUUUUGAAACAUC ..(((((...((((.((....(((((....))))).((.(((((((...............)))))))....))....))--))))..((((.......)))))))))............ ( -35.46) >DroEre_CAF1 1403 114 - 1 AUUUGGCAUGCGUAGCCAAAAGUGCUUAUCAGCACUUGAUUCGGCACAA----AAAAACGCUGCCAAAGCAGCGAAUCAG--UACGAAGUGCUCAGUUUACGCGGCAAUUUUGAAACAUC .....((.((((((.....(((((((....)))))))((((.(((((..----.....((((((....))))))..((..--...)).))))).)))))))))))).............. ( -35.60) >DroYak_CAF1 1380 116 - 1 AUUUGCCAUGCGUAGCCAAAAGUGCUUAUCAGCACUCGAUACGGCACAA----AAAACUGCUGGCAAAGCAGCGAAUCGGUGUACGAAGUGCUCAGUUUACGCGGCAAUUUUGAAACAUC ..(((((..((((((.(...((((((....))))))......(((((..----....(((((.....)))))....(((.....))).)))))..).)))))))))))............ ( -35.20) >consensus AUUUGCCAUGCGUAGCCAAAAGUGCUUAUCAGCACACGAUUCGGCACAA____AAAACUGCUGCCAAAGCAGCGAAACGG__UACGAAGUGCUCAAUUUACGCGGCAAUUUUGAAACAUC ..(((((..(((((.......(((((....)))))..((((.(((((...........((((((....))))))...((.....))..))))).))))))))))))))............ (-27.92 = -29.08 + 1.16)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:11:21 2006