
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 4,794,248 – 4,794,392 |
| Length | 144 |
| Max. P | 0.986874 |

| Location | 4,794,248 – 4,794,364 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 79.17 |
| Mean single sequence MFE | -26.05 |
| Consensus MFE | -15.71 |
| Energy contribution | -16.21 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.69 |
| Structure conservation index | 0.60 |
| SVM decision value | 2.06 |
| SVM RNA-class probability | 0.986874 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
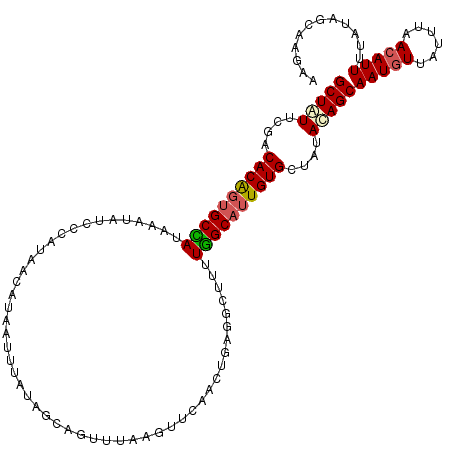
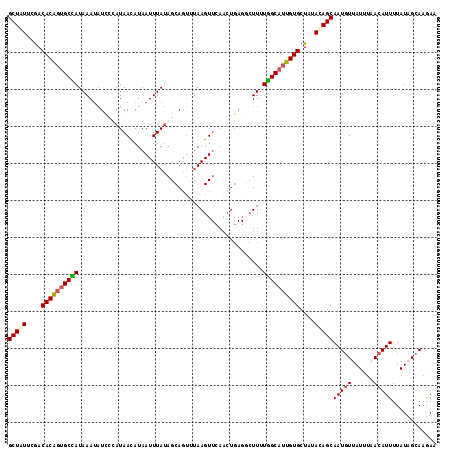
>2L_DroMel_CAF1 4794248 116 + 22407834 GCUCUUCGACACAGUGCUAUAAAAAUCACAUAACAUAAUUUAUAGCAGUGUAAGUUGAACUGAGGCUUUUGGCAUUGUGCUAUACAGCAAUGUUAUUUUACAUUUUAUAGCAAGAA .((((((((((((.(((((((((...............))))))))).)))..))))))..)))((.....))....(((((((....(((((......))))).))))))).... ( -28.86) >DroSec_CAF1 242335 116 + 1 GCUGUUCGACACAGUGCCAUAAAUAUUCCAUAACAUAAUUUAUAGCAGUUUAAGUUCAACUGAGGCUUUUGGCAUUGUGCUAUACAGCAAUGUUAUUUAACAUUUUAUCGCAAGAA (((((....(((((((((((((((.............))))))..(((((.......)))))........)))))))))....)))))((((((....))))))...((....)). ( -30.42) >DroSim_CAF1 219570 105 + 1 GCUAUUCGACACAGUGCCAUAAAUAUUGUAUAACAUAAUUUAUAGCAGUUUAAGU-----------UUUUGGCAUUGUGCUAUACAGCAAUGUUAUUUAACAUUUUAUCGCAAGAA (((......((((((((((.(((((((((((((......)))).)))))....))-----------)).))))))))))......)))((((((....))))))...((....)). ( -26.70) >DroYak_CAF1 230573 109 + 1 GCUUUUUAACACG--GCUAUAAAUCUCCUAUAAUAUAAUUUAAAGCA----AAGUUCAACUGAGCCUUAUAGCUAUGUGCUGCAAAGCAAU-UUAUUGUACAUUUUAUAGAAAAAU (((((...(((..--(((((((.....((.(((......))).))..----..((((....)))).)))))))..))).....)))))...-........................ ( -18.20) >consensus GCUAUUCGACACAGUGCCAUAAAUAUCCCAUAACAUAAUUUAUAGCAGUUUAAGUUCAACUGAGGCUUUUGGCAUUGUGCUAUACAGCAAUGUUAUUUAACAUUUUAUAGCAAGAA (((((....((((((((((..................................................))))))))))....)))))(((((......)))))............ (-15.71 = -16.21 + 0.50)

| Location | 4,794,284 – 4,794,392 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 83.02 |
| Mean single sequence MFE | -23.15 |
| Consensus MFE | -13.77 |
| Energy contribution | -16.40 |
| Covariance contribution | 2.63 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.552173 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
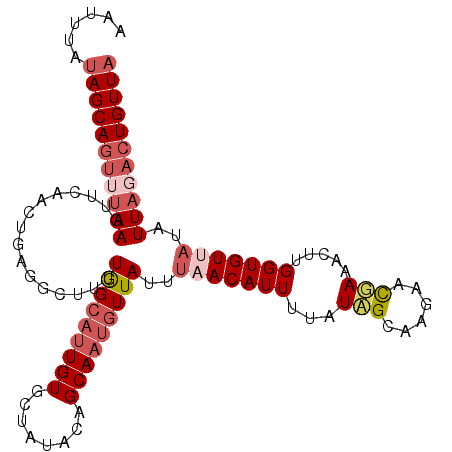
>2L_DroMel_CAF1 4794284 108 + 22407834 AAUUUAUAGCAGUGUAAGUUGAACUGAGGCUUUUGGCAUUGUGCUAUACAGCAAUGUUAUUUUACAUUUUAUAGCAAGAACGAAACUUGGUGUUAUAUUAGACUGUUA ......(((((((.(((.....((..((...((((......(((((((....(((((......))))).)))))))....)))).))..))......))).))))))) ( -26.30) >DroSec_CAF1 242371 108 + 1 AAUUUAUAGCAGUUUAAGUUCAACUGAGGCUUUUGGCAUUGUGCUAUACAGCAAUGUUAUUUAACAUUUUAUCGCAAGAACGAAACUUGGUGUUAUAUUAGACUGUUA ......(((((((((((.....((..(((((..((((.....))))...)))((((((....))))))...((....))......))..))......))))))))))) ( -28.50) >DroSim_CAF1 219606 97 + 1 AAUUUAUAGCAGUUUAAGU-----------UUUUGGCAUUGUGCUAUACAGCAAUGUUAUUUAACAUUUUAUCGCAAGAACGAAACUUGGUGUUAUAUUAGACUGUUA ......(((((((((((..-----------...(((((((((........)))))))))..(((((((...(((......))).....)))))))..))))))))))) ( -24.10) >DroYak_CAF1 230607 103 + 1 AAUUUAAAGCA----AAGUUCAACUGAGCCUUAUAGCUAUGUGCUGCAAAGCAAU-UUAUUGUACAUUUUAUAGAAAAAUUUAAACUUGGUGUUAUAUUAUUCUGUUA .......((((----(.((((....)))).))...)))((((((...(((....)-))...))))))...((((((.(((...(((.....)))..))).)))))).. ( -13.70) >consensus AAUUUAUAGCAGUUUAAGUUCAACUGAGGCUUUUGGCAUUGUGCUAUACAGCAAUGUUAUUUAACAUUUUAUAGCAAGAACGAAACUUGGUGUUAUAUUAGACUGUUA ......(((((((((((................(((((((((........)))))))))..(((((((...(((......))).....)))))))..))))))))))) (-13.77 = -16.40 + 2.63)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:10:19 2006