
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 4,794,080 – 4,794,210 |
| Length | 130 |
| Max. P | 0.995237 |

| Location | 4,794,080 – 4,794,174 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 68.51 |
| Mean single sequence MFE | -25.14 |
| Consensus MFE | -10.78 |
| Energy contribution | -12.10 |
| Covariance contribution | 1.32 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.48 |
| Structure conservation index | 0.43 |
| SVM decision value | 2.56 |
| SVM RNA-class probability | 0.995237 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 4794080 94 + 22407834 CAAUGUUGCUGGCAACAU----------------UGUUGCAGAGGGGAAUUUUCGAGUUCGAAAGUUUCUGGUUCGAUGGAAGGGGGAAUUGA-GAAAAAGCUUGAAUAAA ((((((((....))))))----------------))..((....(((((((((((....)))))))))))..((((((..........)))))-).....))......... ( -24.40) >DroSec_CAF1 242166 95 + 1 CAAUGUUGCUGGCAACAU----------------UGUCGAAGAGGGGAAUUUUCGAGUUCGAAAGUUUCUGAUUCGAAGGAAGGGGGAAAAGGCAAAAAAGCUUGAAUAAA ((((((((....))))))----------------))(((((...(((((((((((....)))))))))))..))))).............((((......))))....... ( -28.40) >DroSim_CAF1 219401 95 + 1 CAAUGUUGCUGGCAACAU----------------UGUCGAAGAGGGGAAUUUUCGAGUUCGAAAGUUUCUGAUUCGUAAGAAGGGGGAAAAGGCAAAAAAGCUUGAAUAAA ((((((((....))))))----------------))(((((((......)))))))((((.....(((((..(((....)))..))))).((((......))))))))... ( -27.30) >DroWil_CAF1 242357 101 + 1 CAAC--CGUUAGCAACAUGCGGGGGUUACUGUUAUUUCGAAAAGCGGGGUUCUAGACUUCGAAAGCUUUUACCUAAAAUC-AGGGGUAAA-------CCAGAUCAAUUAAA ...(--(((.........))))((.(((((.........((((((((((((...))))))....)))))).(((......-)))))))).-------))............ ( -19.50) >DroYak_CAF1 230426 89 + 1 CAAUGUUGCUGGCAACAU----------------GGCCGAAAAGGGGCAUUUUCGAGAUCGAAAACUGCUGCUUCGAAUGAAGGGG-AAAGUGCGAAAAAGCUCCG----- ..((((((....))))))----------------.........(((((.((((((.........(((.((.((((....)))).))-..))).)))))).))))).----- ( -26.10) >consensus CAAUGUUGCUGGCAACAU________________UGUCGAAGAGGGGAAUUUUCGAGUUCGAAAGUUUCUGAUUCGAAGGAAGGGGGAAAAGGCAAAAAAGCUUGAAUAAA ..((((((....))))))..........................(((((((((((....)))))))))))......................................... (-10.78 = -12.10 + 1.32)


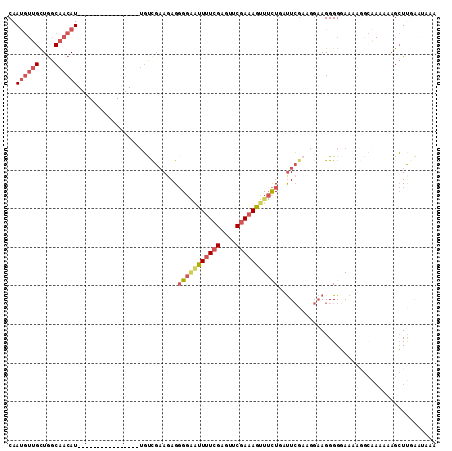
| Location | 4,794,099 – 4,794,210 |
|---|---|
| Length | 111 |
| Sequences | 4 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 74.85 |
| Mean single sequence MFE | -23.78 |
| Consensus MFE | -11.50 |
| Energy contribution | -12.12 |
| Covariance contribution | 0.62 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.48 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.516813 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 4794099 111 + 22407834 GUUGCAGAGGGGAAUUUUCGAGUUCGAAAGUUUCUGGUUCGAUGGAAGGGGGAAUUGA-GAAAAAGCUUGAAUAAACGGUUCAAUUCAAAAUAUACAAAUCUGAUAAACAAG (((.((((.(((((((((((....)))))))))))................(((((((-(...................))))))))............))))...)))... ( -22.91) >DroSec_CAF1 242185 112 + 1 GUCGAAGAGGGGAAUUUUCGAGUUCGAAAGUUUCUGAUUCGAAGGAAGGGGGAAAAGGCAAAAAAGCUUGAAUAAACGGACUAAUACAAAAUAUACAAAUCUAAUAAAUAAG .(((((((......)))))))(((((....(((((..(((....)))..))))).((((......)))).......)))))............................... ( -21.90) >DroSim_CAF1 219420 112 + 1 GUCGAAGAGGGGAAUUUUCGAGUUCGAAAGUUUCUGAUUCGUAAGAAGGGGGAAAAGGCAAAAAAGCUUGAAUAAACGGACUAAUACAAAAUAUACAAAUCUAAUAAAUAAG .(((((((......)))))))(((((....(((((..(((....)))..))))).((((......)))).......)))))............................... ( -22.00) >DroYak_CAF1 230445 98 + 1 GCCGAAAAGGGGCAUUUUCGAGAUCGAAAACUGCUGCUUCGAAUGAAGGGG-AAAGUGCGAAAAAGCUCCG--------AUUGUUGCAAAGCGUGUG-CUGU----AGCAAG ........(((((.((((((.........(((.((.((((....)))).))-..))).)))))).))))).--------.((((((((..((....)-))))----))))). ( -28.30) >consensus GUCGAAGAGGGGAAUUUUCGAGUUCGAAAGUUUCUGAUUCGAAGGAAGGGGGAAAAGGCAAAAAAGCUUGAAUAAACGGACUAAUACAAAAUAUACAAAUCUAAUAAACAAG ..((((...(((((((((((....)))))))))))..))))....................................................................... (-11.50 = -12.12 + 0.62)

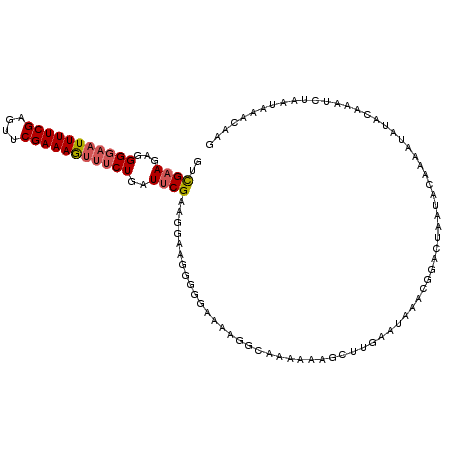
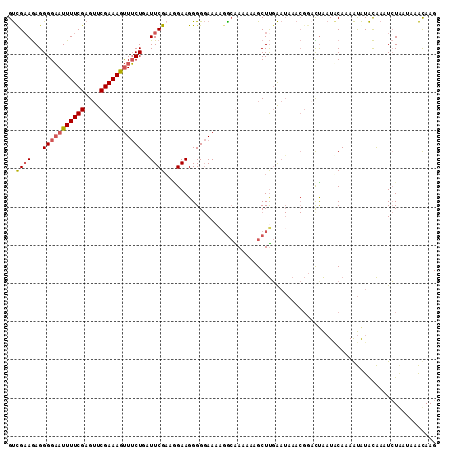
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:10:17 2006