
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 4,772,614 – 4,772,704 |
| Length | 90 |
| Max. P | 0.992543 |

| Location | 4,772,614 – 4,772,704 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 80.19 |
| Mean single sequence MFE | -25.92 |
| Consensus MFE | -15.24 |
| Energy contribution | -17.96 |
| Covariance contribution | 2.72 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.93 |
| Structure conservation index | 0.59 |
| SVM decision value | 2.34 |
| SVM RNA-class probability | 0.992543 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 4772614 90 + 22407834 UGUGUGUGAAGCCGGU-----------UCCACUGGGCACACUGUAAACCAAAAUGAAAAUUGCAAUUGCAAUUGCAAUAAAAUGGCUAAACGCACUUGAGC .((((((..(((((((-----------(.((.((....)).))..)))).........((((((((....)))))))).....))))..))))))...... ( -27.00) >DroSec_CAF1 220931 90 + 1 UGUGCGUGGAGCCGUC-----------UCCACUGGGCACACUGUAAACCAAAAUGAAAAUUGCAAUUGCAAUUGCAAUAAAAUGGCUAAACGCACUUGAGC .((((((..((((((.-----------.....(((............)))........((((((((....))))))))...))))))..))))))...... ( -28.00) >DroSim_CAF1 197886 90 + 1 UGUGCGUGGAGCCGGC-----------UCCACUGGGCACACUGUAAACCAAAAUGAAAAUUGCAAUUGCAAUUGCAAUAAAAUGGCUAAACGCACUUGAGC .((((((..(((((((-----------((....)))).....................((((((((....))))))))....)))))..))))))...... ( -29.70) >DroEre_CAF1 209628 88 + 1 ------UGGAGCCGGUUCCACGGCUCCUCCACU-GGCAGAUUGUAAACCAAAAUGAAAAUUGCA------AUUGCAAUAAAAUGGCUAAACGCACUUGAGC ------.(((((((......))))))).....(-(((..(((((((.....(((....)))...------.)))))))......))))............. ( -23.80) >DroYak_CAF1 207995 88 + 1 ------UGAAGCCGAACCGAACUGCCUUCCACU-GGCAGAUUCUAAACCAAAAUGAAAAUUGCA------AUUGCAAUAAAAUGGCUAAACGCACUUGAGC ------...(((((....((((((((.......-))))).)))...............(((((.------...)))))....))))).............. ( -21.10) >consensus UGUG_GUGGAGCCGGC___________UCCACUGGGCACACUGUAAACCAAAAUGAAAAUUGCAAUUGCAAUUGCAAUAAAAUGGCUAAACGCACUUGAGC .((((((..(((((....................((...........)).........((((((((....))))))))....)))))..))))))...... (-15.24 = -17.96 + 2.72)


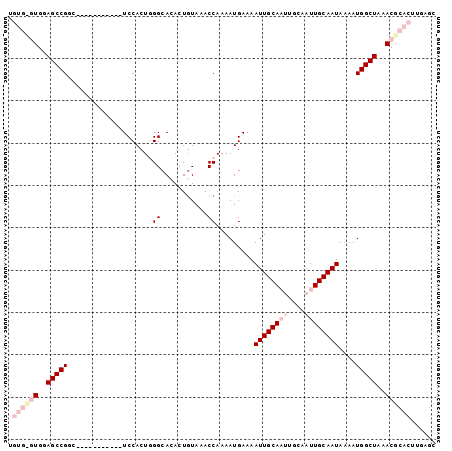
| Location | 4,772,614 – 4,772,704 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 80.19 |
| Mean single sequence MFE | -27.08 |
| Consensus MFE | -15.60 |
| Energy contribution | -16.40 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.57 |
| Structure conservation index | 0.58 |
| SVM decision value | 1.63 |
| SVM RNA-class probability | 0.968725 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 4772614 90 - 22407834 GCUCAAGUGCGUUUAGCCAUUUUAUUGCAAUUGCAAUUGCAAUUUUCAUUUUGGUUUACAGUGUGCCCAGUGGA-----------ACCGGCUUCACACACA ......(((.((..((((.....((((((((....)))))))).........(((((.((.((....)).))))-----------)))))))..)).))). ( -26.30) >DroSec_CAF1 220931 90 - 1 GCUCAAGUGCGUUUAGCCAUUUUAUUGCAAUUGCAAUUGCAAUUUUCAUUUUGGUUUACAGUGUGCCCAGUGGA-----------GACGGCUCCACGCACA ......((((((..((((......(((((((....)))))))((((((((..(((.........))).))))))-----------)).))))..)))))). ( -29.80) >DroSim_CAF1 197886 90 - 1 GCUCAAGUGCGUUUAGCCAUUUUAUUGCAAUUGCAAUUGCAAUUUUCAUUUUGGUUUACAGUGUGCCCAGUGGA-----------GCCGGCUCCACGCACA ......((((((..((((.....((((((((....)))))))).........(((((.((.((....)).))))-----------)))))))..)))))). ( -31.10) >DroEre_CAF1 209628 88 - 1 GCUCAAGUGCGUUUAGCCAUUUUAUUGCAAU------UGCAAUUUUCAUUUUGGUUUACAAUCUGCC-AGUGGAGGAGCCGUGGAACCGGCUCCA------ ((......))......((((...(((((...------.)))))........((((.........)))-))))).(((((((......))))))).------ ( -25.80) >DroYak_CAF1 207995 88 - 1 GCUCAAGUGCGUUUAGCCAUUUUAUUGCAAU------UGCAAUUUUCAUUUUGGUUUAGAAUCUGCC-AGUGGAAGGCAGUUCGGUUCGGCUUCA------ ((......))((..((((.....(((((...------.)))))...............(((.(((((-.......))))))))))))..))....------ ( -22.40) >consensus GCUCAAGUGCGUUUAGCCAUUUUAUUGCAAUUGCAAUUGCAAUUUUCAUUUUGGUUUACAGUGUGCCCAGUGGA___________ACCGGCUCCAC_CACA ((......))....((((.....((((((((....)))))))).(((((...(((.........)))..)))))..............))))......... (-15.60 = -16.40 + 0.80)

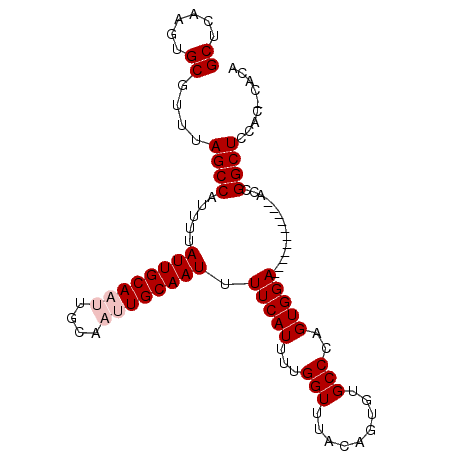
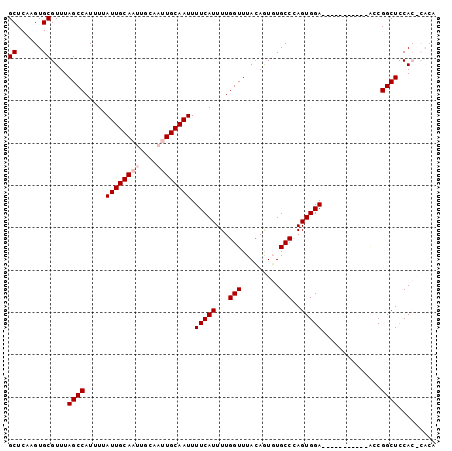
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:10:04 2006